1. Introduction: What is a DNA Fingerprint?
|
A Fingerprint |
Four DNA Fingerprints |
Unless you’re an identical twin, your DNA is completely unique. DNA fingerprinting or DNA profiling involves chemically manipulating DNA to create a pattern that’s unique to every individual (except for identical twins: they’ll share the same DNA fingerprint).
Four DNA fingerprint patterns are shown above. One was made from from DNA collected at a crime scene, and one for each of the three suspects in the crime. Which suspect was at the scene of the crime?
For the past 30 years, DNA fingerprinting has been a mainstay of forensics (crime investigation). It’s also a key tool used in every branch of biological research.
2. RFLPs (Restriction Fragment Length Polymorphisms)
An early method for creating a DNA fingerprint was based on restriction fragments. These segments of DNA are created by bacterial enzymes called restriction endonucleases (frequently referred to as restriction enzymes). These enzymes were discussed in the first tutorial in this module (and you should read about them now if you’re doing this tutorial before that one)
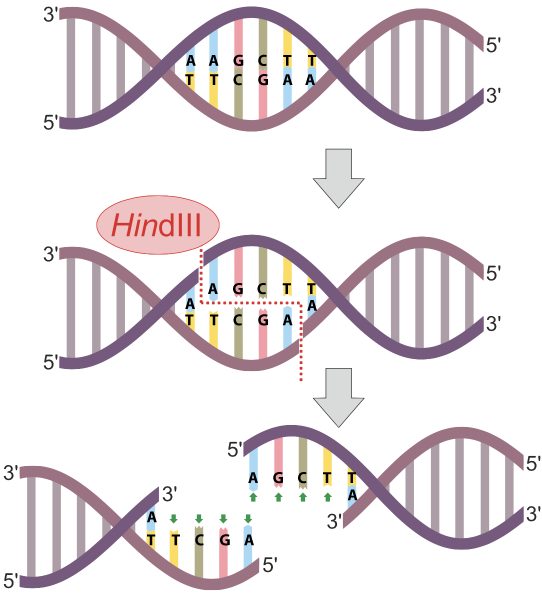
In the diagram at left, the restriction endonuclease HinDIII recognizes the nucleotide sequence AAGCTT, and cuts the DNA between the two As. Note that because that sequence is found on both strands(running in opposite directions), the cut winds up being a double stranded cut, and completely severs the DNA.

If you were to take my DNA and put it into a test tube with restriction enzymes, my DNA would be cut into fragments. Why? Because along the length of the billions of bases in my DNA, there are going to be at least a few instances of restriction site sequences. At these restriction sites, the restriction enzyme will cut up my DNA into what are called restriction fragments.
Now, here’s where the creation of restriction fragments ties into profiling and fingerprinting. A restriction site is a sequence of nucleotide bases. While most of the DNA found in any two humans is the same, there are differences: that’s what makes us genetically distinct. If those differences are within a restriction site, then the number of the restriction fragments created when any two people’s DNA are treated with the same restriction enzyme will also differ, as will the length of those fragments.
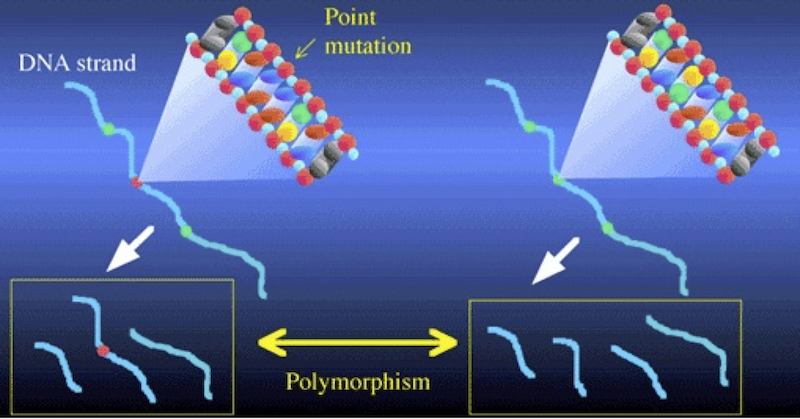
Consider the illustration above. It shows two DNA strands. The sequence on the right has three restriction sites (represented in green). When cut with the corresponding restriction enzyme, that DNA will be cut into four fragments. In the DNA on the left, there’s been a mutation in the second restriction site (shown in red). As a result, the corresponding restriction enzyme won’t recognize that site, and won’t cut the DNA at that point. Consequently, the left sample of DNA is cut into three fragments, while the right sample is cut into four. Note also how in the DNA strand on the left, the combination of the second and third fragments leads one of the fragments to be considerably longer than any of the others.
These differences between the two DNA strands are called polymorphisms. The word polymorphism means “more than one form.” And the basic idea behind this kind of analysis is that differences in restriction sites will result in restriction fragment length polymorphisms. A widely used acronym for “restriction fragment length polymorphism” is RFLP. RFLP differences are of two kinds: differences in number of fragments, and differences in the length of the fragments.
3. RFLPs: Checking Understanding
[qwiz random = “false” qrecord_id=”sciencemusicvideosMeister1961-RFLPs: Checking Understanding (HS)”] [h]
Restriction Enzymes, Fragments, and Polymorphisms
[i]
[q] HindIII is a [hangman] enzyme.
[c]IHJlc3RyaWN0aW9u[Qq]
[f]IENvcnJlY3Qh[Qq]
[q] The two yellow boxes below both show pieces of [hangman] that has been cut into restriction [hangman] .
[c]IEROQQ==[Qq]
[f]IEdyZWF0IQ==[Qq]
[c]IGZyYWdtZW50cw==[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[q] In the diagram below, the restriction fragments are indicated by number
[textentry single_char=”true”]
[c]ID I=[Qq]
[f]IEdyZWF0ISBOdW1iZXIgJiM4MjIwOzImIzgyMjE7IGluZGljYXRlcyBkaWZmZXJlbmNlcyDigJQgcG9seW1vcnBoaXNtcyDigJQgaW4gdGhlIGxlbmd0aHMgb2YgdGhlIHJlc3RyaWN0aW9uIGZyYWdtZW50cyBmcm9tIHR3byBkaWZmZXJlbnQgRE5BIHNhbXBsZXMuwqA=[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IFNvcnJ5LCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBSRkxQcyBhcmUgdGhlIGZyYWdtZW50cyB0aGF0IHJlc3VsdCB3aGVuIEROQSBpcyBjdXQgd2l0aCByZXN0cmljdGlvbiBlbnp5bWVzLiBXaGF0JiM4MjE3O3MgdGhlIG9ubHkgdGhpbmcgaW4gdGhpcyBkaWFncmFtIHRoYXQgY291bGQgcmVwcmVzZW50IGZyYWdtZW50cyBvZiBhIGxhcmdlciBwaWVjZSBvZiBETkE/[Qq]
[q] In the diagram below, number 1 represents a [hangman] in a [hangman] site
[c]IG11dGF0aW9u[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[c]IHJlc3RyaWN0aW9u[Qq]
[f]IE5pY2Ugam9iIQ==[Qq]
[x][restart]
[/qwiz]
4. Gel Electrophoresis
In order to see these restriction fragment length polymorphisms, we take the restriction fragments and subject them to a treatment called gel electrophoresis.
Electrophoresis is “the movement of charged particles in a fluid or gel under the influence of an electric field.” The process involves the type of apparatus shown below. A gel (5), often made of the polysaccharide agarose (a seaweed extract), is placed in an electrophoresis chamber (3). The chamber is essentially a plastic basin which holds a buffer solution that conducts electricity (6). The chamber has positive (4) and negative (2) electrodes, which are attached to a power supply (7).
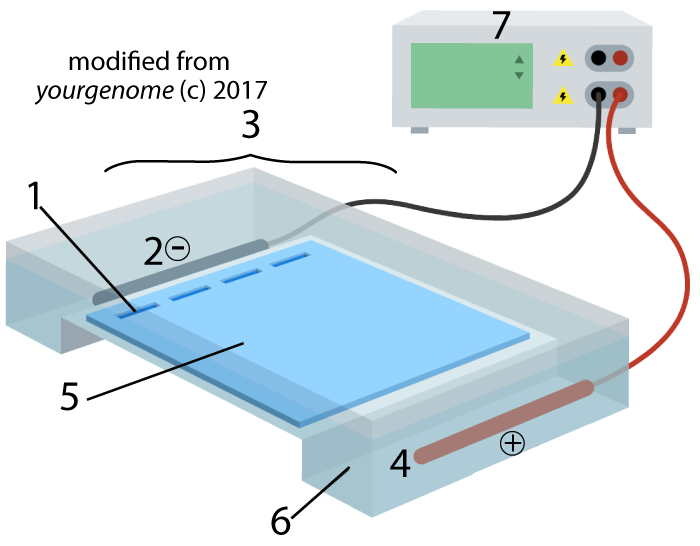
A solution with restriction fragments (also known as a restriction digest) is placed within wells (1), which are indentations in the gel. When you turn the power on, the DNA in the digest will get pushed through the gel, away from the negative electrode (2) and toward the positive electrode (4).
This pushing is related to DNA’s chemistry. DNA’s backbone consists of alternating deoxyribose sugars (the dark blue pentagon with a “D” below) and phosphate groups (the circle with a “P”).
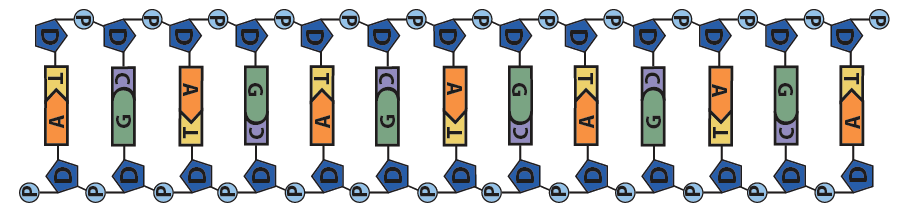
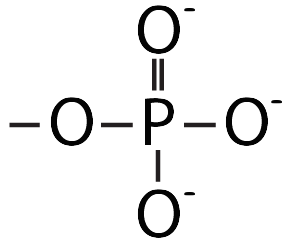
These phosphate groups are negatively charged. As a result, the DNA will be repelled by the negative charge at the negative electrode, and it will be attracted to the positive charge at the positive electrode.
The basic principle of electrophoresis is that small DNA fragments will be impeded by the gel less than large fragments. In other words, large fragments move more slowly than small fragments do, and in a given period of time, small fragments will move further than large ones.
The image below shows what you would see if you put three different samples of DNA into an electrophoresis chamber.
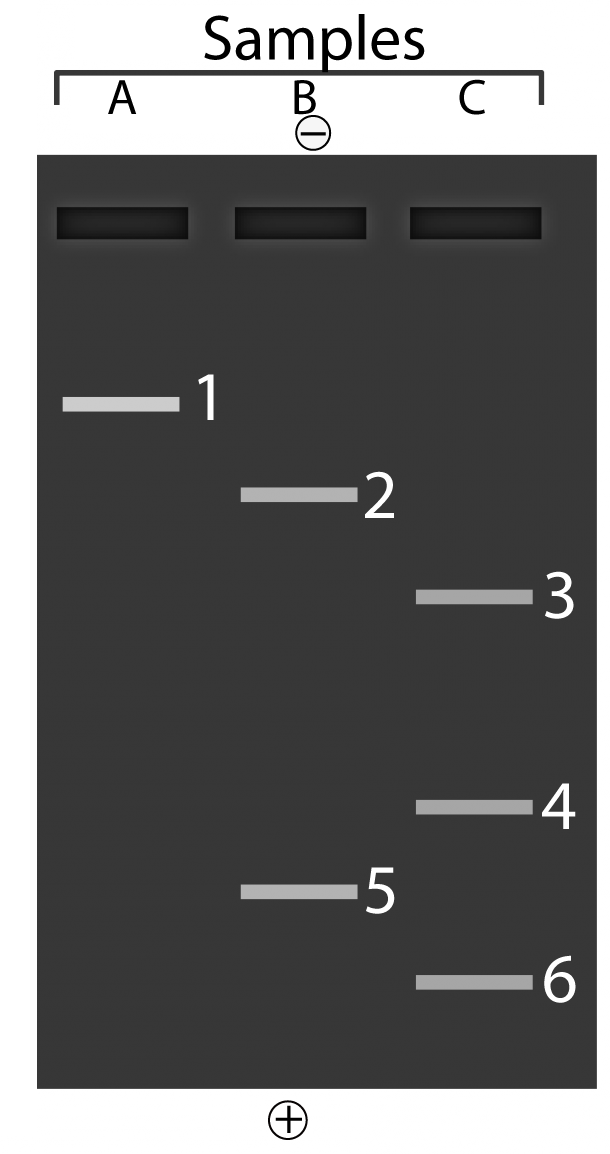 To make sure you’re understanding the explanation above, answer the questions that follow.
To make sure you’re understanding the explanation above, answer the questions that follow.
[qwiz qrecord_id=”sciencemusicvideosMeister1961-DNA Fingerprinting and Electrophoresis (HS)”]
[h]DNA Fingerprinting and gel electrophoresis
[i]
[q]Imagine the you’re a technician in a crime lab for a police department. Where are you going to place your samples of DNA?
[textentry single_char=”true”]
[c]ID E=[Qq]
[f]TmljZS4gJiM4MjIwOzEmIzgyMjE7IHJlcHJlc2VudHMgdGhlICYjODIyMDt3ZWxscyYjODIyMTsgaW4gdGhlIGdlbC4gVGhhdCYjODIxNztzIHdoZXJlIHlvdSYjODIxNztkIHBsYWNlIHRoZSBETkEgc2FtcGxlcy4=[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBZb3UmIzgyMTc7cmUgbG9va2luZyBmb3IgdGhlIHdlbGxzOiBpbmRlbnRhdGlvbnMgaW4gdGhlIGdlbC4gSWYgJiM4MjIwOzUmIzgyMjE7IGlzIHRoZSBnZWwsIHdoYXQgY291bGQgYmUgdGhlIHdlbGxzPw==[Qq]
[q] What’s the smallest fragment in the gel shown below?
[textentry single_char=”true”]
[c]ID Y=[Qq]
[f]TmljZS4gJiM4MjIwOzYmIzgyMjE7IHdvdWxkIGhhdmUgdG8gYmUgdGhlIHNtYWxsZXN0IGZyYWdtZW50Lg==[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[c]ICo=[Qq]
[f]IE5vLiBJbiBnZWwgZWxlY3Ryb3Bob3Jlc2lzLCB0aGUgc21hbGxlc3QgZnJhZ21lbnQgaXMgZ29pbmcgdG8gbW92ZSB0aGUgZmFzdGVzdCwgYW5kIGluIGEgZ2l2ZW4gcGVyaW9kIG9mIHRpbWUsIHRoZSBmYXJ0aGVzdC4=[Qq]
[q] What’s the largest fragment in the gel shown below?
[textentry single_char=”true”]
[c]ID E=[Qq]
[f]RXhjZWxsZW50OiBOdW1iZXIgMSwgd2hpY2ggbW92ZWQgdGhlIHNob3J0ZXN0IGRpc3RhbmNlLCB3b3VsZCBoYXZlIHRoZSBiZSB0aGUgbGFyZ2VzdCBmcmFnbWVudC4=[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[c]ICo=[Qq]
[f]IE5vLiBJbiBnZWwgZWxlY3Ryb3Bob3Jlc2lzLCB0aGUgbGFyZ2VzdCBmcmFnbWVudCBpcyBnb2luZyB0byBtb3ZlIHRoZSBzbG93ZXN0LCBhbmQgaW4gYSBnaXZlbiBwZXJpb2Qgb2YgdGltZSwgdGhlIHNtYWxsZXN0IGRpc3RhbmNlLg==[Qq]
[q] In the diagram below, which letter represents restriction enzymes?
[textentry single_char=”true”]
[c]IG M=[Qq]
[f]IEdvb2QhICYjODIyMDtDJiM4MjIxOyByZXByZXNlbnRzIHJlc3RyaWN0aW9uIGVuenltZXMuwqA=[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IE5vLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50OiByZXN0cmljdGlvbiBlbnp5bWVzIGN1dCBETkEgaW50byBmcmFnbWVudHMu[Qq]
[q] In the diagram below, which letter represents restriction fragments?
[textentry single_char=”true”]
[c]IG Q=[Qq]
[f]IENvcnJlY3QhICYjODIyMDtEJiM4MjIxOyByZXByZXNlbnRzIHJlc3RyaWN0aW9uIGZyYWdtZW50cy4=[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IFNvcnJ5LCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50OiByZXN0cmljdGlvbiBmcmFnbWVudHMgYXJlIHBpZWNlcyBvZiBETkEgdGhhdCBoYXZlIGJlZW4gY3V0IGFwYXJ0IGJ5IHJlc3RyaWN0aW9uIGVuenltZXMu[Qq]
[q] In the diagram below, which letter represents the “gel” in gel electrophoresis?
[textentry single_char=”true”]
[c]IG U=[Qq]
[f]IEdyZWF0ISBMZXR0ZXIgJiM4MjIwO2UmIzgyMjE7IHJlcHJlc2VudHMgdGhlIGdlbC7CoA==[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IE5vLCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBMZXR0ZXIgRCByZXByZXNlbnRzIHJlc3RyaWN0aW9uIGZyYWdtZW50cy4gR2VsIGVsZWN0cm9waG9yZXNpcyBzZXBhcmF0ZXMgRE5BIGZyYWdtZW50cyBieSBzaXplLiBUaGVzZSBmcmFnbWVudHMgYXJlIGdvaW5nIGludG8gX19fX18/[Qq]
[q] In the diagram below, which letter represents the position of the smallest DNA fragments from electrophoresis.
[textentry single_char=”true”]
[c]IG c=[Qq]
[f]IEV4Y2VsbGVudCEgVGhlIHNtYWxsZXN0IGZyYWdtZW50cyB3b3VsZCBtb3ZlIHRoZSBmdXJ0aGVzdCwgYW5kIHdvdWxkIHdpbmQgdXAgYXQgJiM4MjIwO2cuJiM4MjIxOw==[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IFNvcnJ5LCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBEdXJpbmcgZWxlY3Ryb3Bob3Jlc2lzLCB0aGUgc21hbGxlc3QgZnJhZ21lbnRzIG1vdmUgdGhlIGZ1cnRoZXN0Lg==[Qq]
[q] In the diagram below, the DNA fingerprint is represented by
[textentry single_char=”true”]
[c]IG g=[Qq]
[f]IEV4Y2VsbGVudCEgJiM4MjIwO0gmIzgyMjE7IHNob3dzIHRoZSBzb3J0ZWQgZnJhZ21lbnRzLiBUb2dldGhlciwgdGhlIHBhdHRlcm4gdGhleSBmb3JtIG1ha2VzIHVzIGEgRE5BIGZpbmdlcnByaW50LsKg[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IFNvcnJ5LCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBHZWwgZWxlY3Ryb3Bob3Jlc2lzIGlzIGFib3V0IHNvcnRpbmcgdGhlIEROQSBmcmFnbWVudHMgdG8gY3JlYXRlIGEgcGF0dGVybi4gV2hpY2ggcGFydCBvZiB0aGlzIGRpYWdyYW0gY291bGQgcmVwcmVzZW50IHRoaXMgcGF0dGVybi4=[Qq]
[/qwiz]
5. DNA Fingerprinting: Final Thoughts

That’s it for DNA fingerprinting at the high school level. The actual DNA fingerprints used in courtrooms around the world today use a far more elaborate method, first developed by Sir Alec Jeffries in 1984. If you want to learn about it, please follow this link to our AP Bio/College level tutorial about DNA Fingerprinting.
Before leaving, however, consider this. While it might seem like the main use of DNA profiling is to convict criminals, it’s also been used to exonerate people who were wrongly convicted of crimes, often after these people had served years behind bars. The Innocence Project has freed more than 350 wrongly convicted people by using DNA. That includes 20 individuals who were on death row. Click the preceding link to learn more about this group’s work.
Links
- If you’re interested in trying more AP Bio/College Level Biotechnology tutorials, try the preceding link. Tutorials include DNA sequencing, or CRISPR-Cas9, the revolutionary gene editing system.
- Otherwise, choose another topic from the menus above.


