1. All Clades can be taxa…but some (commonly used) taxa are not clades
Any clade (a group derived from a common ancestor) could be a taxon. For example, there’s a name for the clade shown below.
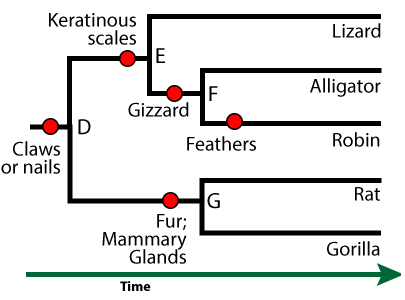
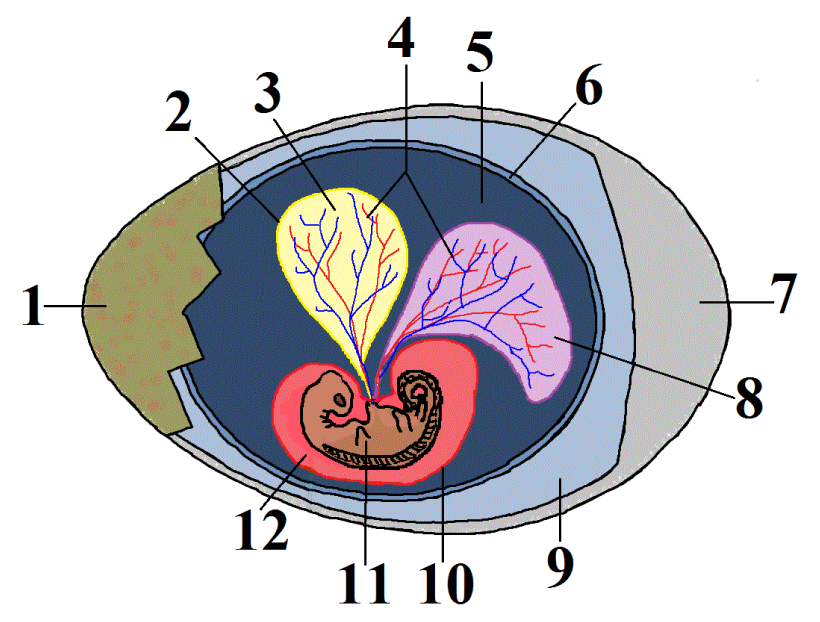
This group is called the amniotes. In addition to possessing the shared derived trait of claws or nails, the females of these species produce a complex egg with an internal, fluid-filled sac that surrounds and nourishes an embryo (shown at 11 at left). This watery internal environment is called an amnion (12). It’s surrounded by an amniotic sac (10).
Note that human beings, like all mammals (including the rat and gorilla in the phylogenetic tree above), are amniotes. Unlike reptiles and birds, mammal eggs, including the amniotic sac and other structures, are kept inside a female’s body.
You’ve probably learned that both alligators and lizards are classified as reptiles. But the taxon we call “reptiles” is not a clade. Remember: a clade includes a common ancestor and all of its descendants.
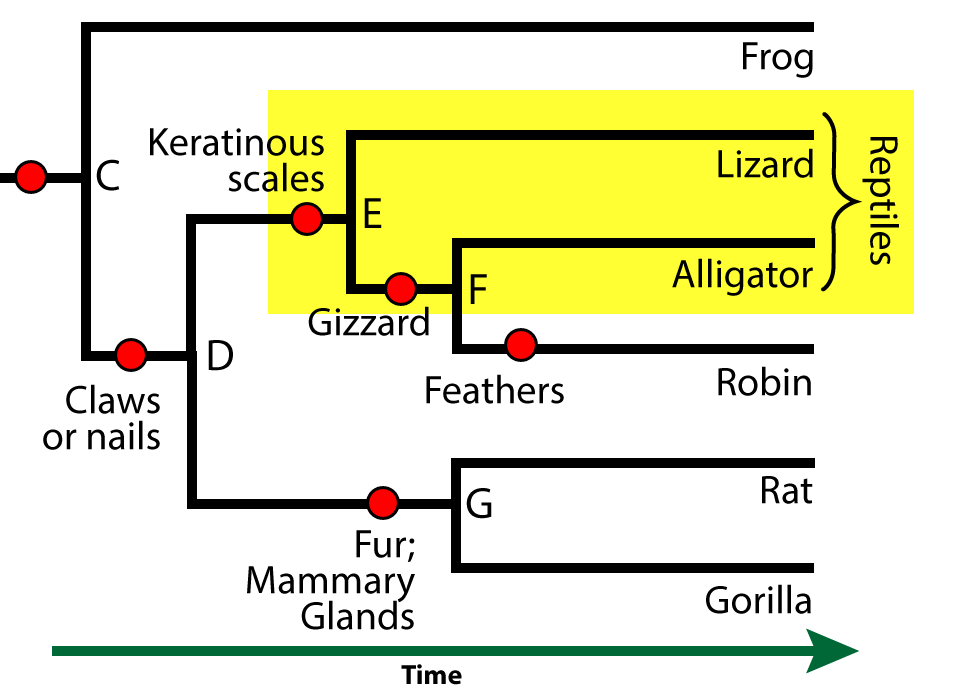 “Reptiles” consists of lizards and alligators (along with other reptiles that aren’t shown here, like snakes), but it excludes birds (with whom the lizards and alligators share a common ancestor).
“Reptiles” consists of lizards and alligators (along with other reptiles that aren’t shown here, like snakes), but it excludes birds (with whom the lizards and alligators share a common ancestor).
Is there a taxon that includes lizards, alligators, and birds? There is one, but it’s not commonly known. It’s called sauropsida, and if you’re interested, you can read more about it in this article on Wikipedia.
Take a look at the groups shown in the cladograms below. Note that all four have the same branching pattern. What’s different are the shaded branches, which indicate different taxa. Some are clades, and some aren’t.
[qwiz qrecord_id=”sciencemusicvideosMeister1961-Is it a clade (HS)”]
[h] Is it a clade?
[q labels = “top”]Label the four trees below. If you’re not sure, use trial and error.
[l]Clade
[f*] Good!
[fx] No. Please try again.
[l]Not a clade
[f*] Good!
[fx] No. Please try again.
[q] Taxa 1 and 2 are clades because they include a common ancestor and all of its [hangman]
[c]IGRlc2NlbmRhbnRz[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[q] Taxon 3 is not a clade because species “c” has a different [hangman] than species “d” and “e.”
[c]IGFuY2VzdG9y[Qq]
[f]IEdvb2Qh[Qq]
[q] The reason why taxon 4 is not a clade is that this taxon is missing one of the [hangman].
[c]IGRlc2NlbmRhbnRz[Qq]
[f]IEdyZWF0IQ==[Qq]
[q multiple_choice=”true”] Based on the discussion above, reptiles would best be represented by which diagram below?
[c]IDEg[Qq][c]IDIg[Qq][c]IDMg[Qq][c]ID Q=
Cg==[Qq][f]IE5vLiAxIGlzIGEgcGVyZmVjdGx5IGdvb2QgY2xhZGUuIFJlcHRpbGVzIGFyZSBub3QgYSBjbGFkZSAoYmVjYXVzZSB0aGV5IGV4Y2x1ZGUgYmlyZHMpLiBXaGljaCBkaWFncmFtIHNob3dzIGFuIGluY29tcGxldGUgY2xhZGU/[Qq]
[f]IE5vLiAyIGlzIGEgcGVyZmVjdGx5IGdvb2QgY2xhZGUuIFJlcHRpbGVzIGFyZSA=bm90IGEgY2xhZGUgKGJlY2F1c2UgdGhleSBleGNsdWRlIGJpcmRzKS4gV2hpY2ggZGlhZ3JhbSBzaG93cyBhbiBpbmNvbXBsZXRlIGNsYWRlPw==[Qq]
[f]IE5vLiBUaGUgcHJvYmxlbSB3aXRoICYjODIyMDszJiM4MjIxOyBpcyB0aGF0IGl0JiM4MjE3O3MgYSB0YXhvbiB3aXRoIG1vcmUgdGhhbiBvbmUgYW5jZXN0b3IuIFJlcHRpbGVzIGFyZSBub3QgYSBjbGFkZSBiZWNhdXNlIHRoZXkgZXhjbHVkZSBvbmUgb2YgdGhlIGRlc2NlbmRhbnRzIChiaXJkcykuIFdoaWNoIGRpYWdyYW0gc2hvd3MgYW4gaW5jb21wbGV0ZSBjbGFkZT8=[Qq]
[f]IEV4Y2VsbGVudC4gJiM4MjIwOzQmIzgyMjE7IGlzIG5vdCBhIGNsYWRlIGluIHRoZSBzYW1lIHdheSB0aGF0IHJlcHRpbGVzIGFyZSBub3QgYSBjbGFkZS4gSXQmIzgyMTc7cyBhIGdyb3VwIHdpdGggYSBjb21tb24gYW5jZXN0b3IsIGJ1dCBub3QgYWxsIG9mIHRoZSBkZXNjZW5kYW50cy4gSW4gdGhlIHNhbWUgd2F5LCB0aGUgcmVwdGlsZSB0YXhvbiBsZWF2ZXMgb3V0IGJpcmRzLg==[Qq]
[/qwiz]
2. Phylogeny and “Molecular Clocks”
In all cladograms and phylogenetic trees, there’s an underlying assumption of the passage of time.
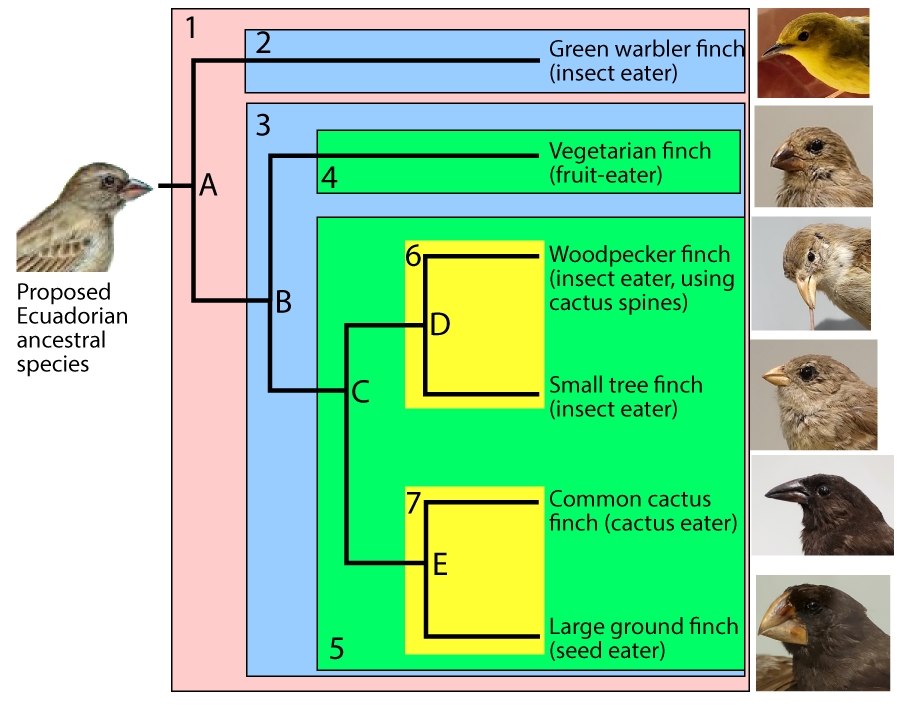
For example, in the phylogenetic tree above, you can assume that clade E emerged more recently than clade C, which emerged more recently than clade B. But there’s no explicit scale.
In other phylogenetic trees, the branch lengths are proportional to the passage of time.
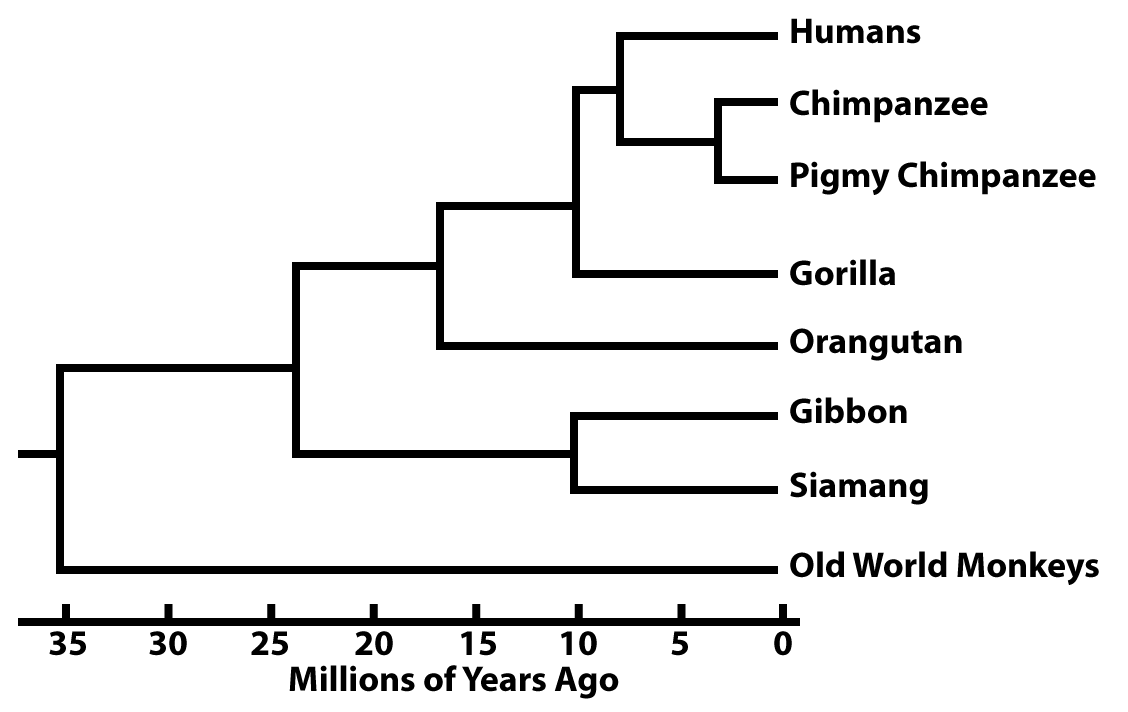
These branch points can be dated from fossil evidence. But increasingly they’re dated based on what are called molecular clocks.
The basic idea behind molecular clocks is that for any specific protein or nucleic acid, the rate of change caused by accumulated mutations is constant over time. If you can calibrate the amount of change in a gene or protein to when two species split apart (a node in the tree), then you can
- Determine a rate of change in that gene or protein,
- Use that rate to determine when other species split in the tree split apart.
Here’s an imaginary example. Imagine a specific sequence of DNA that’s 100 nucleotides long, and it’s shared by all of the primates shown above. In Gibbons and Siamangs, that stretch of DNA is identical except for 10 mutations. Based on fossil evidence, we learn that the split between Gibbons and Siamangs occurred 10 million years ago.
That means that the rate of change in that nucleotide sequence is
- 10 mutations/10 million years, or
- one mutation/million years.
If we find that there are six nucleotide differences in the same stretch of DNA that’s shared by humans and chimps, then a little algebra tells us that humans and chimps would have had a common ancestor about 6 million years ago.
[qwiz qrecord_id=”sciencemusicvideosMeister1961-Molecular Clocks (HS)”]
[h] Molecular Clock Examples
[i]A widely used and highly conserved protein that’s used to determine divergence dates is hemoglobin, the oxygen-carrying protein found in the red blood cells of almost all vertebrates. Hemoglobin consists of two pairs of protein chains. Two of these chains consist of 141 amino acids, and two have 146 amino acids.
The rate of mutation in hemoglobin has been calibrated with changes in the fossil record, allowing scientists to produce the graph below.
[q multiple_choice=”true”] There are 62 amino acid differences between human and frog hemoglobin. Use the graph below to determine when humans and frogs most recently shared a common ancestor.
[c]IDEwMCBtaWxsaW9uIHllYXJz[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgaG93IHRvIHByb2NlZWQuIElkZW50aWZ5IDYyIG9uIHRoZSBZLWF4aXMuIE5vdyBmaW5kIHdoZXJlIDYyIGludGVyY2VwdHMgdGhlIG1vbGVjdWxhciBjbG9jayBsaW5lLCBhbmQgdGhlbiBmaW5kIHRoZSBjb3JyZXNwb25kaW5nIHRpbWUgb24gdGhlIFgtYXhpcy4=[Qq]
[c]IDI1MCBtaWxsaW9uIHllYXJz[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgaG93IHRvIHByb2NlZWQuIElkZW50aWZ5IDYyIG9uIHRoZSBZLWF4aXMuIE5vdyBmaW5kIHdoZXJlIDYyIGludGVyY2VwdHMgdGhlIG1vbGVjdWxhciBjbG9jayBsaW5lLCBhbmQgdGhlbiBmaW5kIHRoZSBjb3JyZXNwb25kaW5nIHRpbWUgb24gdGhlIFgtYXhpcy4=[Qq]
[c]IDM1MCBtaWxs aW9uIHllYXJz[Qq]
[f]RXhjZWxsZW50LiBVc2luZyB0aGUgaGVtb2dsb2JpbiBtb2xlY3VsYXIgY2xvY2ssIHlvdSBjYW4gZXN0aW1hdGUgdGhhdCB0aGUgbGFzdCBjb21tb24gYW5jZXN0b3Igb2YgaHVtYW5zIGFuZCBmcm9ncyBleGlzdGVkIGFib3V0IDM1MCBtaWxsaW9uIHllYXJzIGFnby4=[Qq]
[c]IDUwMCBtaWxsaW9uIHllYXJz[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgaG93IHRvIHByb2NlZWQuIElkZW50aWZ5IDYyIG9uIHRoZSBZIGF4aXMuIE5vdyBmaW5kIHdoZXJlIDYyIGludGVyY2VwdHMgdGhlIG1vbGVjdWxhciBjbG9jayBsaW5lLCBhbmQgdGhlbiBmaW5kIHRoZSBjb3JyZXNwb25kaW5nIHRpbWUgb24gdGhlIFgtYXhpcy4=[Qq]
[q multiple_choice=”true”] Cytochrome c is also used as a molecular clock.
Humans and fruit flies have 29 differences in the amino acids making up their cytochrome c. Based on the graph below, which of the following is closest to the time when humans and fruit flies last shared a common ancestor?
[c]IDEwMCBtaWxsaW9uIHllYXJz[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgaG93IHRvIHByb2NlZWQuIElkZW50aWZ5IDI5IG9uIHRoZSBZIGF4aXMuIE5vdyBmaW5kIHdoZXJlIDI5IGludGVyY2VwdHMgdGhlIG1vbGVjdWxhciBjbG9jayBsaW5lLCBhbmQgdGhlbiBmaW5kIHRoZSBjb3JyZXNwb25kaW5nIHRpbWUgb24gdGhlIFgtYXhpcy4=[Qq]
[c]IDI1MCBtaWxsaW9uIHllYXJz[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgaG93IHRvIHByb2NlZWQuIElkZW50aWZ5IDI5IG9uIHRoZSBZIGF4aXMuIE5vdyBmaW5kIHdoZXJlIDI5IGludGVyY2VwdHMgdGhlIG1vbGVjdWxhciBjbG9jayBsaW5lLCBhbmQgdGhlbiBmaW5kIHRoZSBjb3JyZXNwb25kaW5nIHRpbWUgb24gdGhlIFgtYXhpcy4=[Qq]
[c]IDQwMCBtaWxsaW9uIHllYXJz[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgaG93IHRvIHByb2NlZWQuIElkZW50aWZ5IDI5IG9uIHRoZSBZIGF4aXMuIE5vdyBmaW5kIHdoZXJlIDI5IGludGVyY2VwdHMgdGhlIG1vbGVjdWxhciBjbG9jayBsaW5lLCBhbmQgdGhlbiBmaW5kIHRoZSBjb3JyZXNwb25kaW5nIHRpbWUgb24gdGhlIFgtYXhpcy4=[Qq]
[c]IDUwMCBtaWxs aW9uIHllYXJz[Qq]
[f]IEV4Y2VsbGVudC4gSWYgdGhlcmUgYXJlIDI5IGFtaW5vIGFjaWQgZGlmZmVyZW5jZXMgYmV0d2VlbiBodW1hbiBhbmQgZnJ1aXQgZmx5IGN5dG9jaHJvbWUgYywgdGhlbiB0aGVpciBkaXZlcmdlbmNlIGRhdGUgc2hvdWxkIGJlIGFib3V0IDUwMCBtaWxsaW9uIHllYXJzIGFnbw==[Qq]
[/qwiz]
3. Phylogenetic Trees, Checking Understanding
This module on classification and phylogenetic trees has covered
- Why we need to classify living things.
- How we name species using binomial nomenclature.
- The classification categories used to classify species into taxa above the species level.
- The three domains
- How a key goal of taxonomy is to create taxa that are clades.
- How to analyze lineages, nodes, and common ancestors within phylogenetic trees.
- How to use phylogenetic trees and cladograms to determine how closely related taxa are.
- How to use shared derived features to create phylogenetic trees and cladograms.
- The difference between shared derived features and ancestral features.
- The types of evidence used to construct phylogenetic trees: from anatomical/morphological or molecular (from proteins and nucleic acids).
- How data from DNA and proteins can be used to construct molecular clocks.
If you get a question right, good for you! If you get it wrong, take in the hint that’s provided, and you’ll get another chance. When you’re ready, click start.
[qwiz qrecord_id=”sciencemusicvideosMeister1961-Classification and Phylogenetic Trees: Checking Understanding (HS)”]
[h] Phylogenetic Trees: Checking Understanding
[i]
[q multiple_choice=”true”] In the phylogenetic tree below, the shared derived trait of clade “F” is (a)
[c]IEtlcmF0aW5vdXMgc2NhbGVz[Qq]
[f]IE5vLiBGb3IgY2xhZGUgJiM4MjIwO0YsJiM4MjIxOyBrZXJhdGlub3VzIHNjYWxlcyB3b3VsZCBiZSBhbiBhbmNlc3RyYWwgZmVhdHVyZS4gQSBzaGFyZWQgZGVyaXZlZCBmZWF0dXJlIGlzIG9uZSB0aGF0IHNldHMgdGhlIG1lbWJlcnMgb2YgYSBjbGFkZSBhcGFydCBmcm9tIG90aGVyIGNsYWRlcy4=[Qq]
[c]IEdpen phcmQ=[Qq]
[f]IEV4Y2VsbGVudC4gQSBnaXp6YXJkIHNldHMgdGhlIG1lbWJlcnMgb2YgYSBjbGFkZSAmIzgyMjA7RiYjODIyMTsgYXBhcnQgZnJvbSBvdGhlciBjbGFkZXMsIG1ha2luZyBpdCB0aGVpciBzaGFyZWQgZGVyaXZlZCBmZWF0dXJlLg==[Qq]
[c]IEZlYXRoZXJz[Qq]
[f]IE5vLiBPbmx5IHNvbWUgbWVtYmVycyBvZiBjbGFkZSAmIzgyMjA7ZiYjODIyMTsgaGF2ZSBmZWF0aGVycy4gQSBzaGFyZWQgZGVyaXZlZCBmZWF0dXJlIGlzIG9uZSB0aGF0IGlzIHNoYXJlZCBieSBhbGwgbWVtYmVycyBvZiBhIGNsYWRlLCBhbmQgd2hpY2ggc2V0cyB0aGUgbWVtYmVycyBvZiBhIGNsYWRlIGFwYXJ0IGZyb20gb3RoZXIgY2xhZGVzLiBGaW5kIGEgZmVhdHVyZSB0aGF0IGZpdHMgdGhvc2UgcmVxdWlyZW1lbnRzLg==[Qq]
[q multiple_choice=”true”] In the phylogenetic tree below, an ancestral trait for clade “F” would be
[c]IEtlcmF0aW5v dXMgc2NhbGVz[Qq]
[f]IEV4Y2VsbGVudC4gRm9yIGNsYWRlICYjODIyMDtGLCYjODIyMTsgJiM4MjIwO2tlcmF0aW5vdXMgc2NhbGVzJiM4MjIxOyBpcyBhbiBhbmNlc3RyYWwgZmVhdHVyZS4gSXQgd2FzIGluaGVyaXRlZCBmcm9tIGEgY29tbW9uIGFuY2VzdG9yLCBidXQgbm90IGZyb20gYSBjbG9zZSBlbm91Z2ggYW5jZXN0b3IgdG8gZGlzdGluZ3Vpc2ggdGhpcyBjbGFkZSBmcm9tIG90aGVyIGNsYWRlcy4=[Qq]
[c]IEdpenphcmQ=[Qq]
[f]IE5vLiBBIGdpenphcmQgc2V0cyB0aGUgbWVtYmVycyBvZiBhIGNsYWRlICYjODIyMDtGJiM4MjIxOyBhcGFydCBmcm9tIG90aGVyIGNsYWRlcywgbWFraW5nIGl0IGEgc2hhcmVkIGRlcml2ZWQgZmVhdHVyZS4gQW4gYW5jZXN0cmFsIHRyYWl0IGlzIGFsc28gaW5oZXJpdGVkIGZyb20gYSBjb21tb24gYW5jZXN0b3IsIGJ1dCBpdCYjODIxNztzIG5vdCBhIHRyYWl0IHRoYXQgZGlzdGluZ3Vpc2hlcyB0aGlzIGNsYWRlIGZyb20gb3RoZXIgY2xhZGVzLg==[Qq]
[c]IEZlYXRoZXJz[Qq]
[f]IE5vLiBPbmx5IHNvbWUgbWVtYmVycyBvZiBjbGFkZSAmIzgyMjA7ZiYjODIyMTsgaGF2ZSBmZWF0aGVycywgbWFraW5nIHRoYXQgZmVhdHVyZSBuZWl0aGVyIGEgc2hhcmVkIGRlcml2ZWQgdHJhaXQgbm9yIGFuIGFuY2VzdHJhbCB0cmFpdC4=
Cg==[Qq]
[q] Which of the trees below depicts a taxon has more than one ancestor?
[textentry single_char=”true”]
[c]ID M=[Qq]
[f]IE5pY2Ugam9iISAmIzgyMjA7MyYjODIyMTtoYXMgbW9yZSB0aGFuIG9uZSBhbmNlc3Rvci4=[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IE5vLCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBJbiB0YXhhIDEgYW5kIDIsIHRoZXJlJiM4MjE3O3MgYSBzaW5nbGUgbm9kZSB0aGF0IGxlYWRzIHRvIGFsbCBvZiB0aGUgZGVzY2VuZGFudHMgd2l0aGluIHRoYXQgY2xhZGUuIFdoaWNoIGRpYWdyYW0gc2hvd3MgbW9yZSB0aGFuIG9uZSBub2RlIGJlbmVhdGggdGhlIGRlc2NlbmRhbnRzPw==[Qq]
[q] Which of the trees below depicts a taxon that’s an incomplete clade?
[textentry single_char=”true”]
[c]ID Q=[Qq]
[f]IEdvb2Qgam9iISAmIzgyMjA7NCYjODIyMTsgaXMgYW4gaW5jb21wbGV0ZSBjbGFkZSAoYmVjYXVzZSBpdCYjODIxNztzIG1pc3Npbmcgb25lIG9mIGl0cyBkZXNjZW5kYW50cyku[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IE5vLCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50OiBBIGxvb2sgZm9yIGEgdGF4b24gaW4gd2hpY2ggdGhlIG1lbWJlcnMgc2hhcmUgdGhlIHNhbWUgY29tbW9uIGFuY2VzdG9yLCBidXQgbm90IGFsbCBvZiB0aGUgZGVzY2VuZGFudHMgYXJlIHBhcnQgb2YgdGhlIGNsYWRlLg==[Qq]
[q] If a taxon consists of the ancestor and all of its descendants, then it can be described in one word: a [hangman].
[c]IGNsYWRl[Qq]
[f]IENvcnJlY3Qh[Qq]
[q] The problem with a taxon that included only alligators and lizards is that this taxon would not be a [hangman]. That’s because its common ancestor at letter E was also the ancestor of birds
[c]IGNsYWRl[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[q] A key principle of evolutionary classification is the more closely related two taxa are, the more recently they shared a [hangman] [hangman].
[c]IGNvbW1vbg==[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[c]IGFuY2VzdG9y[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[q] In the diagram below, the most recent common ancestor of A and B would be
[textentry single_char=”true”]
[c]ID E=[Qq]
[f]IEdyZWF0ISAmIzgyMjA7MSYjODIyMTsgaXMgdGhlIG1vc3QgcmVjZW50IGNvbW1vbiBhbmNlc3RvciBvZiBBIGFuZCBCLg==[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IE5vLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBGaW5kIHRoZSBub2RlIHRoYXQgbW9zdCBpbW1lZGlhdGVseSBjb25uZWN0cyAmIzgyMjA7QSYjODIyMTsgYW5kICYjODIyMDtCLiYjODIyMTsgVGhhdCYjODIxNztzIHRoZWlyIG1vc3QgcmVjZW50IGNvbW1vbiBhbmNlc3Rvci7CoA==[Qq]
[q] In the diagram below, the most recent common ancestor of A, B, C, D, and E would be
[textentry single_char=”true”]
[c]ID I=[Qq]
[f]IEF3ZXNvbWUhICYjODIyMDsyJiM4MjIxOyBpcyB0aGUgbW9zdCByZWNlbnQgY29tbW9uIGFuY2VzdG9yIG9mIEEsIEIsIEMsIEQsIGFuZCBFLg==[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IE5vLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBNb3ZlIGJhY2sgaW4gdGltZSBmcm9tIEEsIEIsIEMsIEQsIGFuZCBFIGFuZCBmaW5kIGEgcG9pbnQgd2hlcmUgYWxsIHRoZWlyIGxpbmVhZ2VzIGNvbnZlcmdlLiBUaGF0JiM4MjE3O3MgdGhlaXIgY29tbW9uIGFuY2VzdG9yLsKg[Qq]
[q] If you made up a taxon that included A, B, and C, that taxon would not be a [hangman].
[c]IGNsYWRl[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[q multiple_choice=”true”] According to the diagram below, which of the following statements is true?
[c]IEh1bWFucyBldm9sdmVkIGZyb20gT2xkIFdvcmxkIG1vbmtleXM=[Qq]
[f]IE5vLiAmIzgyMjA7RXZvbHZlZCBmcm9tJiM4MjIxOyBtZWFucywgbW9yZSBvciBsZXNzLCAmIzgyMjA7YXJlIGRlc2NlbmRlZCBmcm9tLiYjODIyMTsgSHVtYW5zIGFuZCBPbGQgV29ybGQgTW9ua2V5cyBhcmUgZXZvbHV0aW9uYXJ5IA==Y291c2lucw==LiBZb3VyIGNvdXNpbnMgYXJlbiYjODIxNzt0IHlvdXIgYW5jZXN0b3JzLg==[Qq]
[c]IEh1bWFucyBhbmQgbmV3IHdvcmxkIG1vbmtl eXMgYmVsb25nIHRvIHRoZSBzYW1lIGNsYWRl[Qq]
[f]IFRoYXQmIzgyMTc7cyBjb3JyZWN0LiBIdW1hbnMgYW5kIG5ldyB3b3JsZCBtb25rZXlzIHNoYXJlIGEgY29tbW9uIGFuY2VzdG9yIGFuZCBhcmUgdGhlcmVmb3JlIG1lbWJlcnMgb2YgdGhlIHNhbWUgY2xhZGUu[Qq]
[q] The study of evolutionary history and relationships is known as [hangman]. (hint: it begins with a “p”).
[c]IHBoeWxvZ2VueQ==[Qq]
[f]IEdyZWF0IQ==[Qq]
[q] A group that consists of a common ancestor and all of its descendants is a [hangman]
[c]IGNsYWRl[Qq]
[f]IEdvb2Qh[Qq]
[q] The two word name for every species, such a Homo sapiens or Bos taurus, is known as a [hangman].
[c]IGJpbm9taWFs[Qq]
[f]IEdvb2Qh[Qq]
[q] In a species name like Turdus migratorius (the American robin), Turdus denotes the [hangman], while migratorius indicates the [hangman].
[c]IGdlbnVz[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[c]IHNwZWNpZXM=[Qq]
[f]IENvcnJlY3Qh[Qq]
[q] Of the classification categories Kingdom, class, and family, the most general one is [hangman], while the most exclusive/specific one is [hangman]
[c]IGtpbmdkb20=[Qq]
[c]ZmFtaWx5[Qq]
[q multiple_choice=”true”] Any named group of organisms is a
[c]IHRh eG9u[Qq]
[f]IENvcnJlY3Qh[Qq]
[c]IHNwZWNpZXM=[Qq]
[f]IE5vLiBXaGlsZSBhIHNwZWNpZXMgaXMgYSB0YXhvbiwgbm90IGFsbCB0YXhhIGFyZSBzcGVjaWVzLsKg[Qq]
[c]IGNsYWRl[Qq]
[f]IE5vLiBNYW55IG5hbWVkIGdyb3VwcyBvZiBvcmdhbmlzbXMgKGxpa2UgJiM4MjIwO3dhdGVyZm93bCwmIzgyMjE7ICYjODIyMDttYXJpbmUgbWFtbWFscywmIzgyMjE7IG9yICYjODIyMDtyZXB0aWxlcyYjODIyMTspIGFyZSBub3QgY2xhZGVzLsKg[Qq]
[q] Letters “A,” “B,” and “C” are points where [hangman] events occurred. They’re called [hangman].
[c]IHNwZWNpYXRpb24=[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[c]IG5vZGVz[Qq]
[f]IEV4Y2VsbGVudCE=[Qq]
[q multiple_choice=”true”] In the phylogenetic tree below, an ancestral trait for clade “F” would be
[c]IEtlcmF0aW5v dXMgc2NhbGVz[Qq]
[f]IEV4Y2VsbGVudC4gRm9yIGNsYWRlICYjODIyMDtGLCYjODIyMTsga2VyYXRpbm91cyBzY2FsZXMgaXMgYW4gYW5jZXN0cmFsIGZlYXR1cmUuIEl0IHdhcyBpbmhlcml0ZWQgZnJvbSBhIGNvbW1vbiBhbmNlc3RvciwgYnV0IG5vdCBmcm9tIGEgY2xvc2UgZW5vdWdoIGFuY2VzdG9yIHRvIGRpc3Rpbmd1aXNoIHRoaXMgY2xhZGUgZnJvbSBvdGhlciBjbGFkZXMu[Qq]
[c]IEdpenphcmQ=[Qq]
[f]IE5vLiBBIGdpenphcmQgc2V0cyB0aGUgbWVtYmVycyBvZiBhIGNsYWRlICYjODIyMDtGJiM4MjIxOyBhcGFydCBmcm9tIG90aGVyIGNsYWRlcywgbWFraW5nIGl0IGEgc2hhcmVkIGRlcml2ZWQgZmVhdHVyZS4gQW4gYW5jZXN0cmFsIHRyYWl0IGlzIGFsc28gaW5oZXJpdGVkIGZyb20gYSBjb21tb24gYW5jZXN0b3IsIGJ1dCBpdCYjODIxNztzIG5vdCBhIHRyYWl0IHRoYXQgZGlzdGluZ3Vpc2hlcyB0aGlzIGNsYWRlIGZyb20gb3RoZXIgY2xhZGVzLg==[Qq]
[c]IEZlYXRoZXJz[Qq]
[f]IE5vLsKgT25seSBzb21lIG1lbWJlcnMgb2YgY2xhZGUgJiM4MjIwO2YmIzgyMjE7IGhhdmUgZmVhdGhlcnMsIG1ha2luZyB0aGF0IGZlYXR1cmUgbmVpdGhlciBhIHNoYXJlZCBkZXJpdmVkIHRyYWl0IG5vciBhbiBhbmNlc3RyYWwgdHJhaXQu[Qq]
[q] The basic idea behind molecular clocks is that for any specific protein or nucleic acid, the rate of change caused by accumulated [hangman] is constant over time.
[c]IG11dGF0aW9ucw==[Qq]
[f]IEdyZWF0IQ==[Qq]
[q] Because mutations accumulate in proteins or nucleic acids at a relatively constant rate, you can infer that mice share a more recent common ancestor with [hangman] and gorillas than they do with chickens.
[c]IG1vbmtleXM=[Qq]
[f]IEdvb2Qh[Qq]
[q] In the diagram below, the shared derived trait of clade G is indicated by number
[textentry single_char=”true”]
[c]ID Q=[Qq]
[f]IEV4Y2VsbGVudC4gJiM4MjIwOzQmIzgyMjE7IHNob3dzIHRoZSBzaGFyZWQgZGVyaXZlZCB0cmFpdCBvZiByYXRzIGFuZCBnb3JpbGxhcyAobWVtYmVycyBvZiB0aGUgbWFtbWFsIGNsYWRlKQ==[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IE5vLCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBMb29rIGZvciBhIHRyYWl0IHRoYXQmIzgyMTc7cyB1bmlxdWUgZm9yIHRoZSBjbGFkZSBpbiB3aGljaCBib3RoIHJhdHMgYW5kIGdvcmlsbGFzIGFyZSBmb3VuZC4=[Qq]
[q] In the diagram below, the common ancestor of lizards, alligators, robins, rats, and gorillas is at letter
[textentry single_char=”true”]
[c]IE Q=[Qq]
[f]IEV4Y2VsbGVudC4gJiM4MjIwO0QmIzgyMjE7IGlzIHRoZSBjb21tb24gYW5jZXN0b3Igb2YgdGhlIGNsYWRlIHRoYXQgdW5pdGVzIGxpemFyZHMsIGFsbGlnYXRvcnMsIHJvYmlucywgcmF0cywgYW5kIGdvcmlsbGFzLg==[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IE5vLCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBMb29rIGZvbGxvdyB0aGUgbGluZWFnZXMgbGVhZGluZyB0byBsaXphcmRzLCBhbGxpZ2F0b3JzLCByb2JpbnMsIHJhdHMsIGFuZCBnb3JpbGxhcyBiYWNrIGluIHRpbWUgdW50aWwgdGhleSBtZWV0IGluIGEgY29tbW9uIGFuY2VzdG9yLg==[Qq]
[q] In the diagram below, what number indicates the most recent ancestral trait of lizards, alligators, robins, rats, and gorillas?
[textentry single_char=”true”]
[c]ID I=[Qq]
[f]IEV4Y2VsbGVudC4gJiM4MjIwOzImIzgyMjE7IGlzIHRoZSBtb3N0IHJlY2VudCBhbmNlc3RyYWwgdHJhaXQgb2YgdGhlIGNsYWRlIHRoYXQgdW5pdGVzIGxpemFyZHMsIGFsbGlnYXRvcnMsIHJvYmlucywgcmF0cywgYW5kIGdvcmlsbGFzICh0aGUgc2hhcmVkIGRlcml2ZWQgdHJhaXQgaXMgJiM4MjIwOzMuJiM4MjIxOw==[Qq]
[c]IEVudGVyIHdvcmQ=[Qq]
[f]IFNvcnJ5LCB0aGF0JiM4MjE3O3Mgbm90IGNvcnJlY3Qu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBGaW5kIHRoZSBzaGFyZWQgZGVyaXZlZCB0cmFpdCBvciB0aGUgY2xhZGUgdGhhdCB1bml0ZXMgbGl6YXJkcywgYWxsaWdhdG9ycywgcm9iaW5zLCByYXRzLCBhbmQgZ29yaWxsYXMuIFRoZW4gZmluZCB0aGUgdHJhaXQgdGhhdCB3b3VsZCBpbmNsdWRlIHRoZSBvdXRncm91cCBvZiB0aGF0IGNsYWRlICh3aGljaCBpcyB0aGUgZnJvZyku[Qq]
[/qwiz]
What now?
This tutorial ends this high school level module about classification and phylogenetic trees. Use this link to return to the Classification and Phylogenetic Trees Main Menu (HS Level). Or choose another tutorial from the menus above.
