For an overview of everything below, start with this video.
Page Outline
- The Functions of Proteins
- Proteins are Polymers of Amino Acids, Linked by Peptide Bonds
- Amino Acids, Peptide Bonds, and Polypeptide Quiz
- Protein structure
- The Dynamic Nature of Proteins
- Denaturation
- Proteins quiz
1. The Functions of Proteins
Let’s start our study of proteins by considering our hands.

Our fingernails are composed of a protein called keratin. A different form of keratin makes up the outermost layer of your skin. The elasticity of deeper layers of your skin — your skin’s ability to return to its shape after being pinched or stretched — is made possible by a protein called elastin.
Go a bit deeper. Your fingers move because they’re attached to muscles by tendons and ligaments that are composed of a protein called collagen. The muscles that move the bones in your hand are made of proteins such as actin and myosin.
Think about your veins. The red blood cells that flow within them are filled with an oxygen-carrying protein called hemoglobin. If you were to cut your hand and get an infection, your immune system would produce proteins called immunoglobulins to form the antibodies that would beat back the invading bacteria.
When you look at any animal, you’re looking at protein. The same is not true of plants, which are mostly made of carbohydrates. But how did those carbohydrates get there? The enzyme that pulls carbon dioxide out of the air to create carbohydrates during photosynthesis is called Rubisco. Like almost all enzymes, it’s a protein.
Let’s organize our thinking about the various functions of proteins by completing the interactive table below. Use prior knowledge and trial and error (and don’t worry about getting every answer right the first time).
[qwiz qrecord_id=”sciencemusicvideosMeister1961-Functions of Proteins (2.0)”]
[h] Some of the functions of proteins
[q labels = “top”]Proteins:
- Control chemical reactions as ______________.
- Create _____________ like bone, hair, feathers (keratin)
- Fight __________ (antibodies)
- Produce movement (___________)
- ____________ oxygen (hemoglobin in red blood cells)
- Store ____________________ (albumin in egg white)
- Transmit _________ as hormones and neurotransmitters.
PROTEINS ARE THE ___________________ BIOLOGICAL MACROMOLECULES
[l]chemical energy
[f*] Excellent!
[fx] No, that’s not correct. Please try again.
[l]infection
[f*] Correct!
[fx] No. Please try again.
[l]enzymes
[f*] Excellent!
[fx] No, that’s not correct. Please try again.
[l]MOST DIVERSE
[f*] Great!
[fx] No, that’s not correct. Please try again.
[l]muscle
[f*] Correct!
[fx] No. Please try again.
[l]structures
[f*] Excellent!
[fx] No. Please try again.
[l]transport
[f*] Correct!
[fx] No. Please try again.
[l]signals
[f*] Good!
[fx] No. Please try again.
[/qwiz]
2. Proteins are polymers of amino acids, linked by peptide bonds
2a. Amino Acids
Proteins are polymers of amino acids. There are twenty amino acids found in living things. Each one has a three-letter abbreviation: “lys” for lysine, “ser” for serine, etc. All are variations of a common structure.
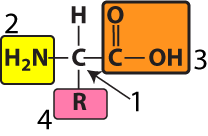
At the center of every amino acid is a central carbon atom (at “1” in the diagram to your left). The central carbon is also referred to as the alpha (α) carbon. This central carbon is flanked by an amino group (at “2”) and a carboxylic acid (at “3”). That’s where the name “amino acid” comes from.
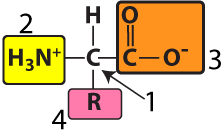
Note that the amino group and the carboxyl group can be shown in both their non-ionized (as shown on the left) and their ionized forms (as shown on the right). You should be able to recognize both.
At “4” is an “R group” or “side chain.” Each of the twenty amino acids has a distinct R group. The R groups are shaded in the diagram below.
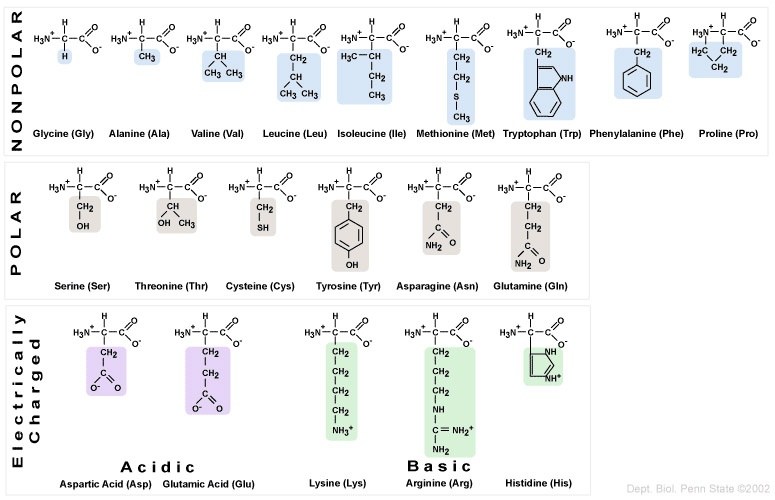
The twenty amino acids can be organized into four categories, based on the chemistry of their side chains. These R groups can be
- Non-polar/hydrophobic
- Polar/hydrophilic
- Acidic (leaving them with a negative charge after they’ve donated a proton to the solution)
- Basic (leaving them with a positive charge after they’ve absorbed a proton from the solution)
Following a few rules, you can easily identify which category an amino acid falls into.
- Non-polar: R groups with methyl functional groups (—CH3), or with fused carbon rings.
- Polar: R groups with hydroxyl (—OH) or sulfhydryl (—SH) functional groups.
- Acidic: R groups with carboxylic acid functional groups.
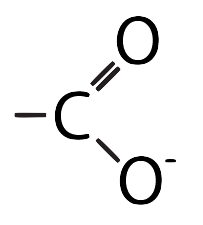
- Basic: R groups with amino functional groups: NH3+ or NH2+ or NH.
This is important because mutations in DNA can result in substitutions of one amino acid for another. If the substituted amino acid has a different chemistry than the original amino acid, the effect on the protein’s structure and function can be significant.
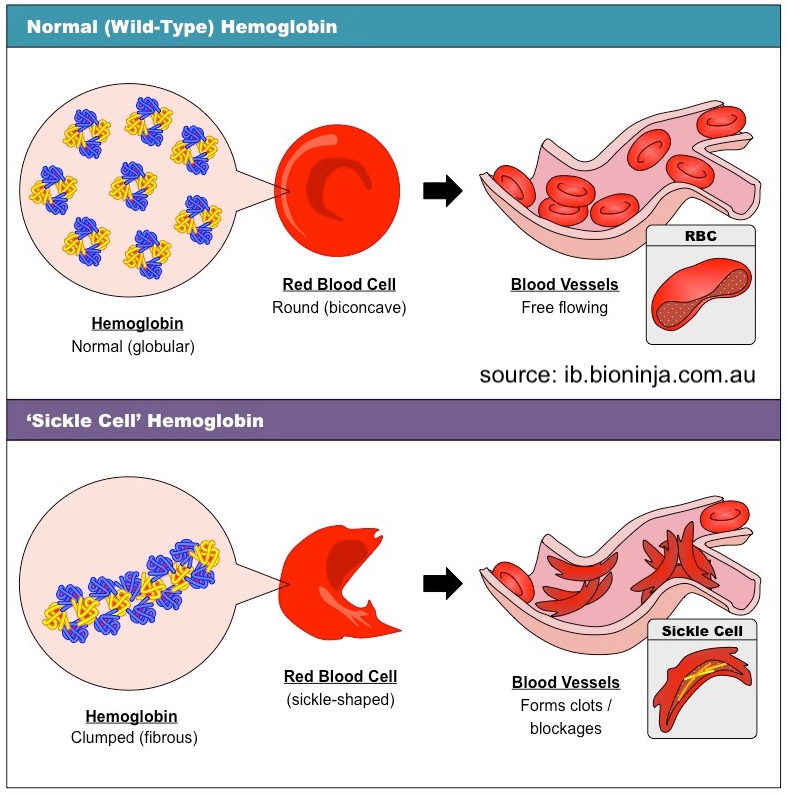 For example, the inherited blood disorder sickle cell anemia is caused by a substitution of glutamic acid for valine in a protein called hemoglobin, which carries oxygen in red blood cells. Glutamic acid is (obviously) an acid, while valine is non-polar.
For example, the inherited blood disorder sickle cell anemia is caused by a substitution of glutamic acid for valine in a protein called hemoglobin, which carries oxygen in red blood cells. Glutamic acid is (obviously) an acid, while valine is non-polar.
The substitution changes hemoglobin’s chemistry so that those mutated hemoglobin molecules, when under low-oxygen conditions, form hydrophobic bonds with one another, creating a fibrous chain as shown in the lower half of the diagram on the left. This causes red blood cells to become sickled instead of smooth. These sickled blood cells form clots and blockages in capillaries (tiny blood vessels), which cause tissue damage and pain.
2b. Peptide Bonds
During the process of protein synthesis, ribosomes (tiny particles within cells, represented by “2” below) link amino acids (“1”) together, following instructions sent to them from the cell’s DNA. The instructions are in the form of a single-stranded nucleic acid called messenger RNA, shown at “3” below.
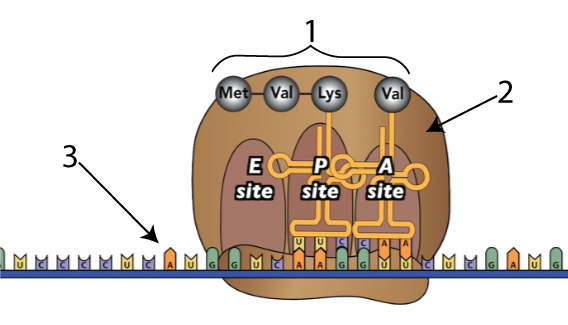
Here’s what happens as ribosomes link amino acids together.
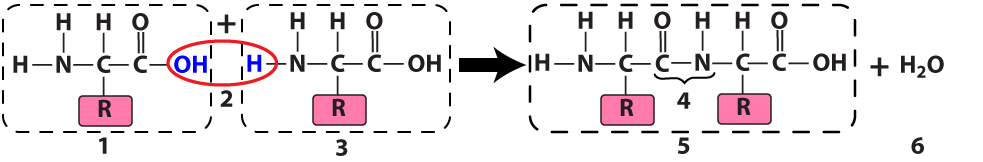
The ribosome connects two amino acids (“1” and “3”) by removing a hydroxyl functional group (an —OH) from one amino acid and a hydrogen atom from another. The result is a dipeptide (“5”): two amino acids linked by a peptide bond (at “4”). Because this is a dehydration synthesis, a water molecule (at “6”) also results from this reaction.
2c. Polypeptides
Add more amino acids and you have a polypeptide (a chain of linked amino acids).
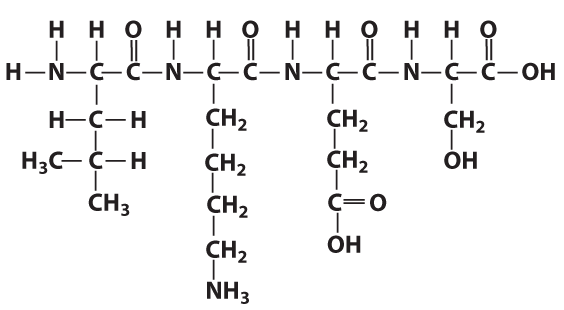
In the same way as the letters in a word can be in any order, the amino acids in a protein can be in any order. As I’ve noted previously, however, the “letters to words = amino acids to protein” analogy is limited. Proteins can be thousands of amino acids long, making the universe of potential proteins much greater than the universe of potential words.
Let’s end this section with a bit of protein terminology (all of which has shown up on recent AP bio exams).
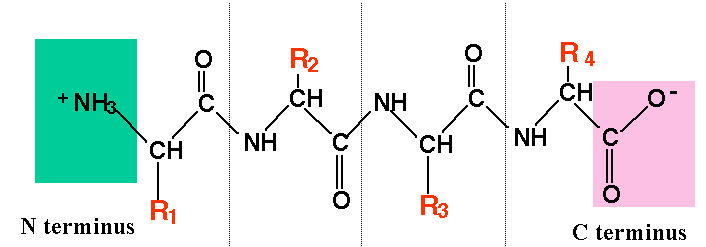 Each of the ends of a protein/polypeptide chain has a specific name. The amino acid at the start of a protein has an exposed amino functional group (an —NH2). That amino acid is called the N terminal amino acid, and that end is called the N terminus. The amino acid at the other end of a protein has an exposed carboxyl group (—COOH), making that the C-terminal amino acid, and that end of the C-terminus. Why is this important? For one thing, it defines the capabilities of protein-digesting enzymes, some of which can only break off amino acids at the N terminus, while others are limited to the C terminus. Second, it constrains what happens during protein synthesis, because ribosomes can only add new amino acids at the carboxyl terminus of a growing polypeptide chain.
Each of the ends of a protein/polypeptide chain has a specific name. The amino acid at the start of a protein has an exposed amino functional group (an —NH2). That amino acid is called the N terminal amino acid, and that end is called the N terminus. The amino acid at the other end of a protein has an exposed carboxyl group (—COOH), making that the C-terminal amino acid, and that end of the C-terminus. Why is this important? For one thing, it defines the capabilities of protein-digesting enzymes, some of which can only break off amino acids at the N terminus, while others are limited to the C terminus. Second, it constrains what happens during protein synthesis, because ribosomes can only add new amino acids at the carboxyl terminus of a growing polypeptide chain.- An amino acid within a polypeptide chain is sometimes referred to as a residue (because it’s what’s left over after the dehydration synthesis reaction that put it there). For example, in a discussion of the mutation in hemoglobin that causes sickle cell anemia, you might read that “the mutation is in the sixth amino acid residue,” as shown below
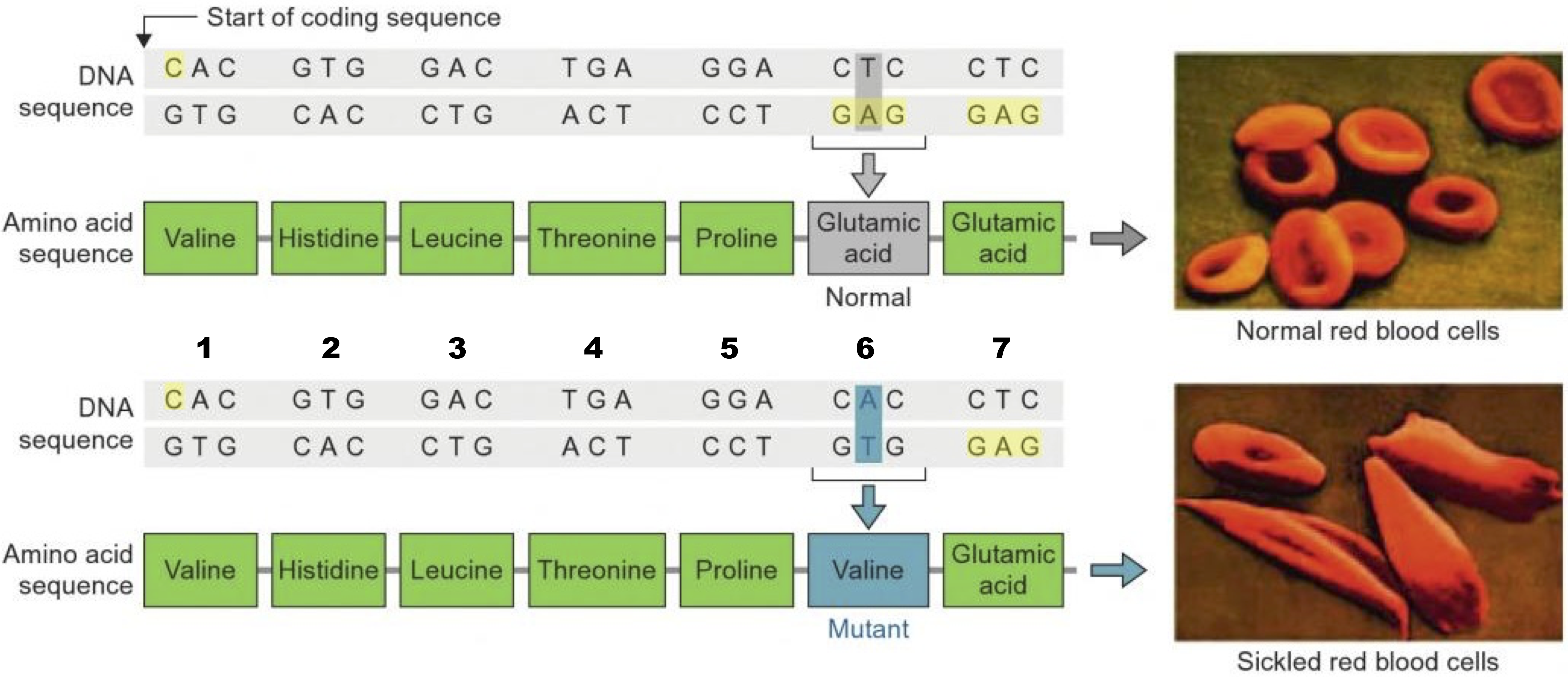
That just means that the mutation is in the sixth amino acid of the chain.
3. Amino Acids, Peptide Bonds, and Polypeptides: Checking Understanding
[qwiz use_dataset=”SMV_biochem_7_amino acids and peptide bonds” dataset_intro=”false” random=”true” qrecord_id=”sciencemusicvideosMeister1961-Amino Acids, Polypeptides (2.0)”]
[h]Amino Acids, Peptide Bonds, and Polypeptides
[i]
[x][restart][/qwiz]
4. Protein Structure
4a. Proteins are Specific and Dynamic
Protein structure has a few remarkable features. The first is specificity of shape. To get an understanding of this, let’s think about our immune system.
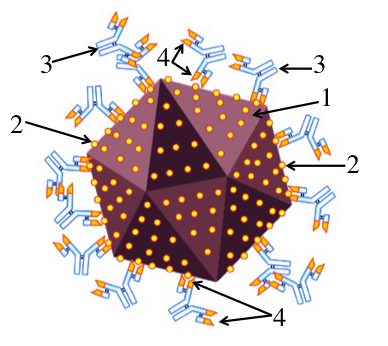
Imagine that you’ve been infected by a virus (“1”). Your immune system counterattacks by producing proteins called antibodies (“3”) that can bind with specific molecules on the surface of the virus. These molecules are called antigens (“2”).
An antibody can bind with an antigen because the tip of the antibody has an antigen-binding site (“4”) whose shape closely matches with, or complements, the shape of the antigen. This binding either directly neutralizes the virus (keeping it, for example, from infecting a cell), or allows other parts of the immune system to neutralize and destroy the virus.
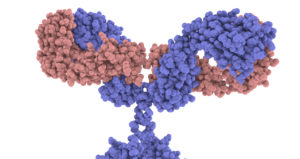
In the diagram above, I drew the antigen as a circle and gave the antigen-binding site on the antibody a complementary shape. But to get a sense of the required specificity of the antigen-binding site, click on the image on the right, from the Garvan Institute of Medical Research. This is a space-filling model, with every atom represented. Note that I’ve set up the image so you can click on it to open an enlarged version of it. When you click, take a close look at the antigen-binding sites on the tips of the arms.
The key point is that this shape, down to the nanometer scale, is specific. It can only do its job (binding with the viral antigen) if it has this exact shape. And this leads to the question: how can protein shape be determined so specifically?
The second feature of protein shape that needs to be explained is that for many proteins, the shape is dynamic. It can change.
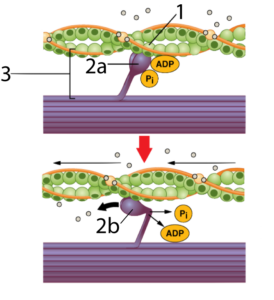 Let’s consider what happens when you contract a muscle. Two interacting proteins are involved: actin (at “1”) and myosin (at “2a” and “2b”). The two interact to form a cross-bridge (at “3”), which is made possible by complementarity in shape between the head of the myosin and a binding site on the actin.
Let’s consider what happens when you contract a muscle. Two interacting proteins are involved: actin (at “1”) and myosin (at “2a” and “2b”). The two interact to form a cross-bridge (at “3”), which is made possible by complementarity in shape between the head of the myosin and a binding site on the actin.
When myosin binds with ATP (the cell’s energy-transfer molecule) the ATP is broken into its lower energy form (ADP and Pi). This enables the myosin to reach and grab onto actin (as shown in “2a”). When the myosin releases ADP and Pi, it again changes shape, bending into a right angle (as shown at “2b”), which pulls on the actin, shortening the muscle fiber.
In other words, the myosin protein is like a lever: provide it with ATP (and meet a few other conditions) and it will bend. Other protein-based structures spin like propellers.
Take three minutes and watch the video below, created by XVIVO for Harvard University.
Pretty much every mind-blowing thing that’s represented involves the dynamic nature of proteins. So the question is what kind of molecular structure allows a protein (like myosin) to change its shape?
4b. Primary Structure
The specificity of protein shape, as well as many proteins’ ability to change shape, emerges from three or four levels of molecular interactions, usually referred to as “levels of structure.” The first level, also called primary structure, is determined genetically.
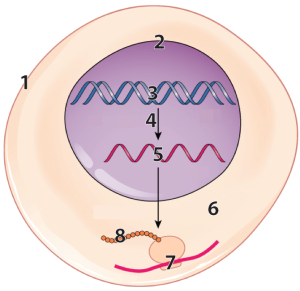
DNA (“3”) in the nucleus (“2”) of our cells has instructions for making RNA (“5”).
That RNA is synthesized through a process called transcription, which is indicated by the arrow at “4.” After transcription, a variety of processes in the nucleus modify the RNA, converting it into messenger RNA (mRNA).
The mRNA leaves the nucleus and enters the cytoplasm (“6”), where ribosomes (“7”) translate the RNA into a polypeptide (“8”).

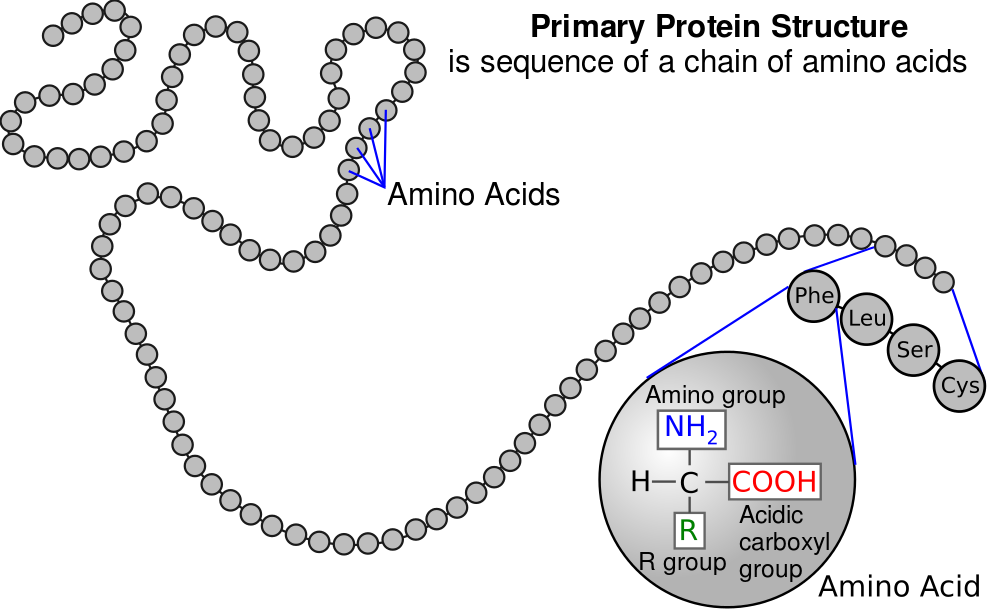
Note that each of the little circles in “8” above is an amino acid residue. Now, connect that image to what you’ve learned about amino acids, peptide bonds, and polypeptides. “8” above is, biochemically, what you see on the right: a linear sequence of amino acid residues in a polypeptide chain.
 So, now we need to explain how we get from the genetically determined linear string of amino acids in a polypeptide chain to a protein’s three-dimensional shape.
So, now we need to explain how we get from the genetically determined linear string of amino acids in a polypeptide chain to a protein’s three-dimensional shape.
4c. Secondary Structure
The next level of interaction involves hydrogen bonds that form between carbonyl functional groups and amino groups within the backbone of a polypeptide. To see this, take a look at the diagram of a polypeptide below:
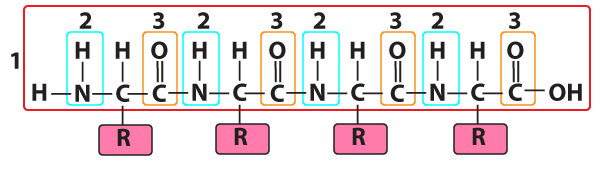
Number “1” is the polypeptide backbone. In the same way that your ribs are attached to your backbone, the side chains/R groups of each amino acid residue are attached to the polypeptide backbone.
The polypeptide was formed by connecting the amino group of one amino acid to the carboxyl group of the next, in a dehydration synthesis reaction that removed an “—H” from the amino group and an “—OH” from the carboxyl group. This leaves behind two recognizable functional groups. At “2” there’s an —NH (still called an “amino group,” despite the loss of a hydrogen atom). At “3” is a carbonyl group. Note that it was a carboxyl group (—COOH), but having lost its terminal —OH, it’s now carbonyl (—C=O).
Both the amino group and the carbonyl group are polar. The amino has a partially positive charge (δ+), and the carbonyl has a partially negative charge (δ–). That means these two functional groups, if they get close enough to one another, can form a hydrogen bond. And while hydrogen bonds are relatively weak (much weaker than the covalent bonds that hold the polypeptide backbone together), they can twist the polypeptide backbone into particular shapes, and stabilize those shapes. The two shapes are called an alpha helix, and a beta-pleated sheet.
4c.1. Alpha Helix
An alpha helix is a corkscrew, stabilized by hydrogen bonds.
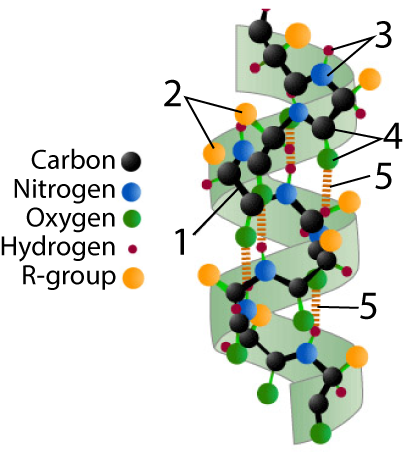
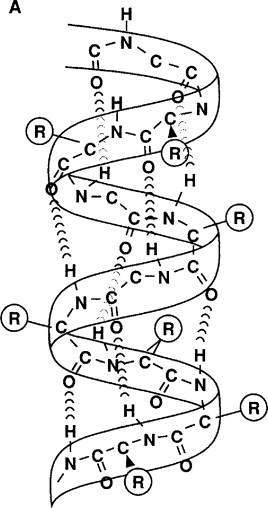
Below, I’ve provided two representations of an alpha helix. The one on the left is a ball and stick model. The one on the right has symbols for each element. As you read below, switch back and forth between the two models.
|
|
|
In the ball-and-stick model, the polypeptide backbone is indicated by “1.” You should be able to notice the repeating pattern of carbon-carbon-nitrogen within the backbone.
R-groups (shown at “2”) hang off of each central carbon. The nitrogen atoms are bonded to hydrogen atoms, forming amino groups (“3”), while the second carbon atom (the one that’s not the central carbon) is bonded to an oxygen, creating a carbonyl group (“4”).
In an alpha helix, hydrogen bonds (shown at “5” form between the carbonyl group and amino groups in amino acid residues that are four amino acids away from one another.
4c.2. Beta Pleated Sheet
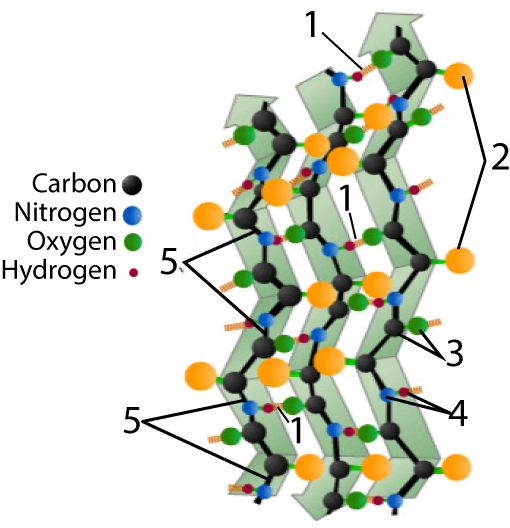
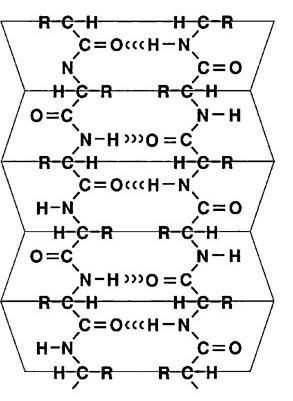
The other secondary structure to know is a beta-pleated sheet.
Again, you’ll find two models below. As you read below, switch back and forth between them.
|
|
|
4d. Tertiary Structure
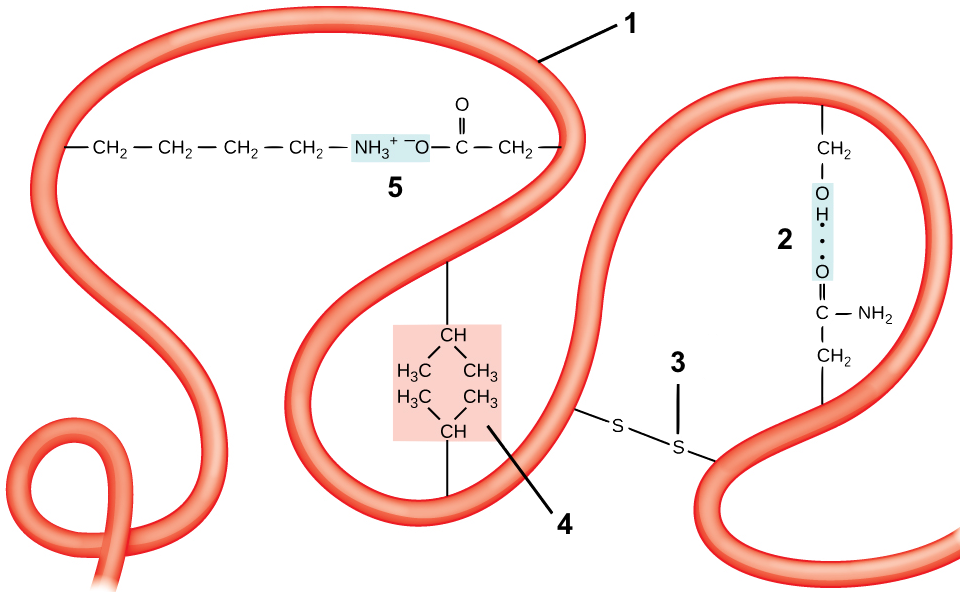
Tertiary structure comes about through interactions between R groups, which cause a polypeptide chain (“1”) to twist and bend. These interactions include
- Hydrogen bonds: if a side chain with a carbonyl group (which has a partially negative charge) gets close to a hydroxyl group or an amino group, a hydrogen bond can form. One such bond between a carbonyl and a hydroxyl is shown at “2.”
- Disulfide Bridges: the amino acid cysteine has a sulfhydryl functional group (—SH) at the end of its side chain. If two of these sulfhydryl groups from cysteines in the same polypeptide chain get close to one another, they can form a disulfide bridge. A disulfide bridge is a covalent bond.
- Hydrophobic interactions: If hydrophobic side chains come into proximity, they can form hydrophobic clusters. These clusters are maintained by two forces. The first is the exclusion of these hydrophobic side chains from the hydrogen bonding between water molecules in the surrounding solution. The second is the weak Van Der Waals bonds that form between these hydrophobic side chains. One such hydrophobic cluster is shown at “4.”
- Ionic bonds: These form when side chains with full positive charges come into proximity with side chains that have full negative charges, as is shown in “5.”
4e. Quaternary Structure
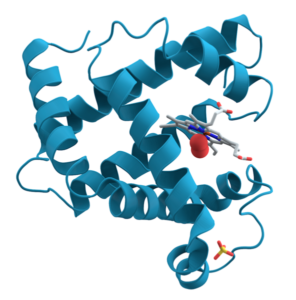
For some proteins, primary, secondary, and tertiary interactions explain the protein’s form and function. Myoglobin, for example, is a protein that stores and releases oxygen within muscle tissue. It consists of 154 amino acids that twist, turn, and coil into the shape shown at the right. As you look, I’m hoping that you’re seeing the alpha helices (and connecting them to secondary structure) and hairpin turns (and thinking that some tertiary bond must be causing the polypeptide chain to turn at such a sharp angle).
In other proteins, there’s a fourth level of structure, called quaternary structure. Quaternary structure involves two or more folded polypeptide chains interacting to form a more complex structure.
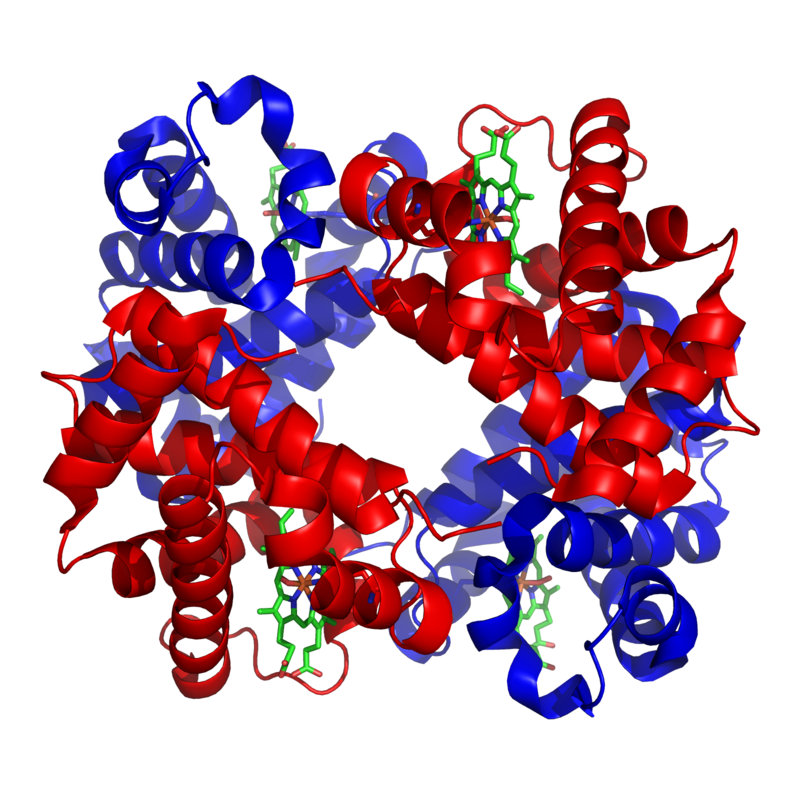
An example of one such quaternary-level protein is hemoglobin. Hemoglobin (which we discussed above in the context of sickle cell disease) is the molecule that carries oxygen in our red blood cells. It consists of four polypeptide chains, two alpha chains (shown in red), and two beta chains (shown in blue).
The bonds between the polypeptide subunits in a quaternary protein can include any of the bonds that appear at the tertiary level: hydrogen bonds, hydrophobic interactions, disulfide bridges, and ionic bonds.
4f. The Dynamic Nature of Proteins
My goal in the sections above was to explain how proteins have such specific shapes. But the second question we wanted to answer about proteins was how they can change their form to act as molecular versions of gates, levers, pulleys, vice grips, propellers, and so on.
 The basic idea is that when proteins bind with other molecules, when their environment changes (through temperature changes or pH changes), or when they are compressed or stretched, the configuration of bonds within that protein can change. Changing the configuration of bonds, in turn, can change the protein’s shape.
The basic idea is that when proteins bind with other molecules, when their environment changes (through temperature changes or pH changes), or when they are compressed or stretched, the configuration of bonds within that protein can change. Changing the configuration of bonds, in turn, can change the protein’s shape.
An alpha helix, for example, is like a molecular spring. A change in another part of the protein can compress that spring, or stretch it out, after which it might return to its original form. Or, if a tertiary bond that caused a hairpin turn in a polypeptide chain is altered, then that chain might straighten out, only to snap back later.
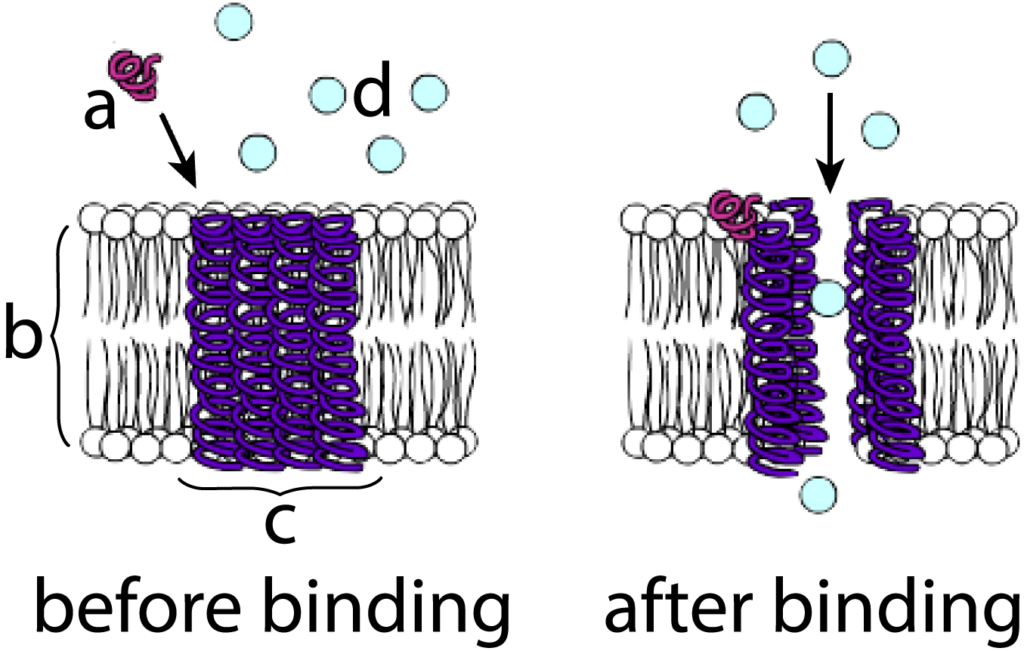
Let’s use the example of a membrane protein that acts as a calcium gate as an example. The diagram at left shows a protein hormone (“a”) diffusing toward the cell membrane (“b”). Embedded within the membrane is a channel protein (“c”). In such membrane channels, alpha helices are often a predominant structure, and you should be able to see several of these helices spanning the membrane.
The binding of the hormone with the channel protein (shown at right), changes the configuration of these alpha-helices so that they move apart from one another. This creates an opening in the membrane where one didn’t exist before. Substances can now diffuse through this opening. When the hormone, which is only loosely bound to the channel protein through hydrogen bonds, dissociates from the channel, the channel will close up, and diffusion will stop.
5. Protein Structure Flashcards
[qdeck bold_text=”false” style=”width: 550px !important; min-height: 450px !important;” random=”true” qrecord_id=”sciencemusicvideosMeister1961-Protein Structure Flashcards”]
[h] Protein Structure Flashcards
[i]
Normal (left) and Mutated Hemoglobin
[start][q json=”true” yy=”4″ unit=”1.Chemistry_of_Life” dataset_id=”AP_Bio_Flashcards_2022|1f1607a7aa110″ question_number=”30″ topic=”1.5-6.Proteins_and_Nucleic_Acids”] What’s the monomer of proteins? Describe that monomer’s structure.
[a] Amino acids are the monomers of proteins. They’re built around a central carbon, attached to an amino group (2), a carboxyl group (3), a hydrogen atom, and a variable R group (4). The R group also called a “side chain” can be polar, non-polar, acidic, or basic. Interactions between amino acids (covered in another card) determine the protein’s three-dimensional shape.
[q json=”true” yy=”4″ unit=”1.Chemistry_of_Life” dataset_id=”AP_Bio_Flashcards_2022|1f0bfd3462d10″ question_number=”31″ topic=”1.5-6.Proteins_and_Nucleic_Acids”] List 6 key functions of proteins. Here’s a hint: Each of the letters in the acronym MESTES refers to a function.
[a] Proteins functions include:
- Motion (as in the proteins actin and myosin, which interact to create muscle tissue);
- Controlling chemical reactions as enzymes;
- Structure (as in the flexible protein collagen or the more fibrous keratin, which makes hair, feathers, and nails;
- Transport (as in hemoglobin, which carries oxygen)
- Energy storage (albumin)
- Transmitting signals (as in protein hormones like insulin or protein neurotransmitters like serotonin).
[q json=”true” yy=”4″ unit=”1.Chemistry_of_Life” question_number=”32″ topic=”1.5-6.Proteins_and_Nucleic_Acids” dataset_id=”AP_Bio_Flashcards_2022|6535dac8b8285″] Describe primary structure.
[a] Primary structure consists of the sequence of amino acids in a polypeptide chain. This sequence is genetically determined.
In the diagram below, each sphere represents one of the linked amino acids making up a protein’s primary structure.
[q json=”true” yy=”4″ unit=”1.Chemistry_of_Life” question_number=”33″ topic=”1.5-6.Proteins_and_Nucleic_Acids” dataset_id=”AP_Bio_Flashcards_2022|65282d02dca85″] Describe secondary structure.
[a] Secondary protein structure involves interactions between carbonyl groups and amino groups in the polypeptide backbone. These interactions can cause a polypeptide to twist into a coiled alpha helix (left), or form a regularly folded structure called a pleated sheet (right).
[q json=”true” yy=”4″ unit=”1.Chemistry_of_Life” question_number=”34″ topic=”1.5-6.Proteins_and_Nucleic_Acids” dataset_id=”AP_Bio_Flashcards_2022|11a98dedfa14a9″] Describe tertiary structure.
[a] Interactions between amino acid side chains (also called R-groups) result in a tertiary structure. These interactions involve hydrogen bonds (2), ionic bonds (5), covalent bonds (3), or hydrophobic clustering (4). The result is a complex, three-dimensional shape.
[q json=”true” yy=”4″ unit=”1.Chemistry_of_Life” question_number=”35″ topic=”1.5-6.Proteins_and_Nucleic_Acids” dataset_id=”AP_Bio_Flashcards_2022|11a8a0713d3ca9″] Describe quaternary structure.
[a] Multiple polypeptides can interact to form a quaternary structure, which is found in proteins such as hemoglobin (shown below), which consists of four, interconnected, polypeptides. The bonds that stabilize a quaternary structure include hydrogen bonds, ionic bonds, and hydrophobic interactions.
[q json=”true” yy=”4″ unit=”1.Chemistry_of_Life” dataset_id=”AP_Bio_Flashcards_2022|1eebf990e3d10″ question_number=”36″ topic=”1.5-6.Proteins_and_Nucleic_Acids”] What’s a C-terminus? An N-terminus?
[a] Every protein has an amino group on one end (the amino terminus), and a carboxyl group on the other end (the carboxyl terminus). This turns out to be biologically significant during both protein synthesis and protein digestion.
[q json=”true” yy=”4″ unit=”1.Chemistry_of_Life” dataset_id=”AP_Bio_Flashcards_2022|1ee1a49c20110″ question_number=”37″ topic=”1.5-6.Proteins_and_Nucleic_Acids”] Describe the structure and function of hemoglobin, and explain the molecular cause of sickle cell disease.
[a] Hemoglobin is a quaternary protein made of four polypeptide chains. It transports oxygen in red blood cells.
People with sickle cell disease inherit a mutation in which the amino acid valine (with a nonpolar side chain) substitutes for glutamic acid (with an acidic side chain). Consequently, when blood becomes deoxygenated, the mutated hemoglobin molecules form hydrophobic bonds with one another, causing them to aggregate into fibers. This reduces hemoglobin’s capacity to carry oxygen and deforms the shape of red blood cells, which become elongated and spiked. This causes these cells to become trapped in capillaries, blocking blood flow and causing pain and tissue damage.
[/qdeck]
6. Denaturation
A protein’s function is based on its structure. As we’ve seen above, the interactions that result in protein structure at the secondary, tertiary, and quaternary levels involve some forces that are relatively weak, such as hydrogen bonds and other Van Der Waals bonds. Other interactions, such as ionic bonds, only occur if a solution has a specific pH. Proteins, in a word, can be delicate, and their function can be affected by changes in their environment. A protein that loses its function because of environmental change —usually heat or pH — is said to be denatured, and the process by which this happens is called denaturation. A quick look at enzymes will show you how this works (you can learn more about enzymes here).
Enzymes are catalysts that speed up chemical reactions in living things. With very few exceptions, enzymes are proteins. Here’s an example of how an enzyme might work to break the disaccharide lactose into two monosaccharides, glucose and galactose
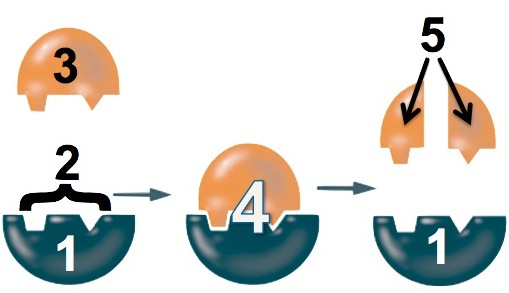
- The enzyme (“1”) binds with the lactose (“3”). The binding is only possible because a specific part of the enzyme, its active site (“2”), has a shape that complements the shape of the lactose.
- While the enzyme is bound to the lactose, the enzyme changes shape to stress the bonds that hold the glucose and galactose monomers together.
- The bond breaks and glucose and galactose are released from the enzyme.
While the details of different enzyme-catalyzed reactions will vary, the key point is that the enzyme can catalyze the reaction only if it can fit together with its substrate (whatever molecule it interacts with). That fit is as specific as the ability of a key to open a lock. Change the key’s shape, and the lock won’t open.
With these ideas in hand, let’s look at the graph below, which shows the relationship between the rate of an enzyme-catalyzed reaction, the percentage of active enzyme molecules, and temperature.
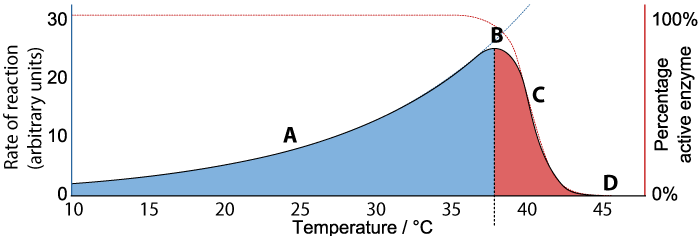
We’ll consider the rate first. As the temperature increases from 10° C to about 37°C (“A”), the rate of the reaction also increases. At about 37°C, the reaction reaches a peak (“B”). As the temperature continues to increase much beyond 37°C, the rate of the reaction declines (“C”) until at 45°C the rate of the reaction falls to zero.
Now consider the “percentage of active enzyme” (the Y-axis on the right). For any temperature between 10° C to just beyond 37°C, the percentage of active enzyme is 100%. But after point “B,” the percentage drops off (“C”) until it reaches zero (“D”).
As I’m sure you’ve surmised, what’s happening is denaturation. Up to a certain temperature, the enzyme can hold its shape and interact with its substrate to bring about the reaction. As temperature increases up to 37°C, the reaction proceeds faster and faster, because higher temperature means more molecular movement and a greater chance that the enzyme will bump into its substrate at the right orientation to bring the reaction about. Beyond 37°C, the kinetic energy in the system starts to disrupt the bonds that are holding the enzyme in its required three-dimensional shape. As this shape starts to change, the enzyme becomes unable to bond with its substrate, and the rate of the reaction (along with the amount of active enzyme) falls to zero.
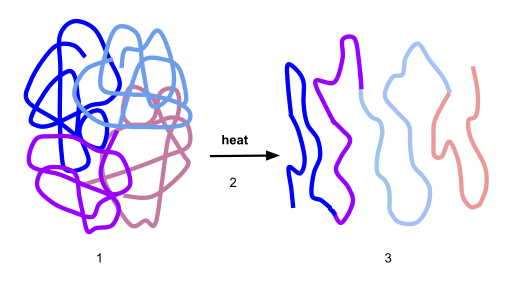 Here’s a visual representation of denaturation. Number “1” shows the protein in its optimal conformation. Heat (“2”) breaks the bonds that stabilize this protein’s shape at the secondary and tertiary levels. As a result, the enzyme unfurls into a polypeptide.
Here’s a visual representation of denaturation. Number “1” shows the protein in its optimal conformation. Heat (“2”) breaks the bonds that stabilize this protein’s shape at the secondary and tertiary levels. As a result, the enzyme unfurls into a polypeptide.
Sometimes, like when you cook an egg and change liquid albumin into solid egg white, the denaturation is irreversible. In other words, you might let your egg white cool down, but it’s not going to liquefy again. On the other hand, some proteins can temporarily be denatured (usually by changes in pH). When the pH is reset to the original level, the amino acids in the polypeptide might be able to interact in a way that restores the protein’s original shape. That process is called renaturation.
7. Proteins Cumulative Quiz
[qwiz use_dataset=”SMV_biochem_8_proteins_cumulative” dataset_intro=”false” random=”true” qrecord_id=”sciencemusicvideosMeister1961-Proteins Cumulative quiz (2.0)”] [h]
Proteins Cumulative Quiz
[i]
[x][restart][/qwiz]
Links
Topic 1.6: Nucleic Acids Overview (the last tutorial in AP Bio Unit 1)